Dense and pleiotropic regulatory information in a developmental enhancer
- PMID: 33057197
- PMCID: PMC8236315
- DOI: 10.1038/s41586-020-2816-5
Dense and pleiotropic regulatory information in a developmental enhancer
Abstract
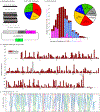
Changes in gene regulation underlie much of phenotypic evolution1. However, our understanding of the potential for regulatory evolution is biased, because most evidence comes from either natural variation or limited experimental perturbations2. Using an automated robotics pipeline, we surveyed an unbiased mutation library for a developmental enhancer in Drosophila melanogaster. We found that almost all mutations altered gene expression and that parameters of gene expression-levels, location, and state-were convolved. The widespread pleiotropic effects of most mutations may constrain the evolvability of developmental enhancers. Consistent with these observations, comparisons of diverse Drosophila larvae revealed apparent biases in the phenotypes influenced by the enhancer. Developmental enhancers may encode a higher density of regulatory information than has been appreciated previously, imposing constraints on regulatory evolution.
Figures









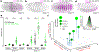

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

