Identification of sequence-specific interactions of the CD44-intracellular domain with RUNX2 in the transcription of matrix metalloprotease-9 in human prostate cancer cells
- PMID: 33062960
- PMCID: PMC7556329
- DOI: 10.20517/cdr.2020.21
Identification of sequence-specific interactions of the CD44-intracellular domain with RUNX2 in the transcription of matrix metalloprotease-9 in human prostate cancer cells
Abstract
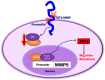
Aim: The Cluster of differentiation 44 (CD44) transmembrane protein is cleaved by γ-secretase, the inhibition of which blocks CD44 cleavage. This study aimed to determine the biological consequence of CD44 cleavage and its potential interaction with Runt-related transcription factor (RUNX2) in a sequence-specific manner in PC3 prostate cancer cells.
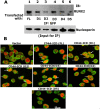
Methods: Using full-length and C-terminal deletion constructs of CD44-ICD (D1-D5) expressed as stable green fluorescent protein-fusion proteins in PC3 cells, we located possible RUNX2-binding sequences.
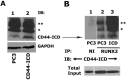
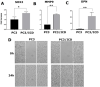
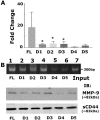
Results: Chromatin immunoprecipitation assays demonstrated that the C-terminal amino acid residues between amino acids 671 and 706 in D1 to D3 constructs were indispensable for sequence-specific binding of RUNX2. This binding was minimal for sequences in the D4 and D5 constructs. Correspondingly, an increase in matrix metalloprotease-9 (MMP-9) expression was observed at the mRNA and protein levels in PC3 cells stably expressing D1-D3 constructs.
Conclusion: These results provide biochemical evidence for the possible sequence-specific CD44-ICD/RUNX2 interaction and its functional relationship to MMP-9 transcription in the promoter region.
Keywords: CD44; CD44-ICD; MMP-9; Prostate cancer; RUNX2; metastasis; tumorigenesis.
Conflict of interest statement
Conflicts of interest All authors declared that there are no conflicts of interest.
Figures



Similar articles
-
Characterization of CD44 intracellular domain interaction with RUNX2 in PC3 human prostate cancer cells.Cell Commun Signal. 2019 Jul 22;17(1):80. doi: 10.1186/s12964-019-0395-6. Cell Commun Signal. 2019. PMID: 31331331 Free PMC article.
-
Integrin αvβ3 and CD44 pathways in metastatic prostate cancer cells support osteoclastogenesis via a Runx2/Smad 5/receptor activator of NF-κB ligand signaling axis.Mol Cancer. 2012 Sep 11;11:66. doi: 10.1186/1476-4598-11-66. Mol Cancer. 2012. PMID: 22966907 Free PMC article.
-
RUNX2 interacts with BRG1 to target CD44 for promoting invasion and migration of colorectal cancer cells.Cancer Cell Int. 2020 Oct 15;20:505. doi: 10.1186/s12935-020-01544-w. eCollection 2020. Cancer Cell Int. 2020. PMID: 33071648 Free PMC article.
-
CD44: A Multifunctional Cell Surface Adhesion Receptor Is a Regulator of Progression and Metastasis of Cancer Cells.Front Cell Dev Biol. 2017 Mar 7;5:18. doi: 10.3389/fcell.2017.00018. eCollection 2017. Front Cell Dev Biol. 2017. PMID: 28326306 Free PMC article. Review.
-
Regulatory roles of Runx2 in metastatic tumor and cancer cell interactions with bone.Cancer Metastasis Rev. 2006 Dec;25(4):589-600. doi: 10.1007/s10555-006-9032-0. Cancer Metastasis Rev. 2006. PMID: 17165130 Review.
Cited by
-
Hyaluronan Functions in Wound Repair That Are Captured to Fuel Breast Cancer Progression.Biomolecules. 2021 Oct 20;11(11):1551. doi: 10.3390/biom11111551. Biomolecules. 2021. PMID: 34827550 Free PMC article. Review.
-
CD44 In Sarcomas: A Comprehensive Review and Future Perspectives.Front Oncol. 2022 Jun 17;12:909450. doi: 10.3389/fonc.2022.909450. eCollection 2022. Front Oncol. 2022. PMID: 35785191 Free PMC article. Review.
-
CD44v8-10 is a marker for malignant traits and a potential driver of bone metastasis in a subpopulation of prostate cancer cells.Cancer Biol Med. 2021 May 20;18(3):788-807. doi: 10.20892/j.issn.2095-3941.2020.0495. Cancer Biol Med. 2021. PMID: 34018387 Free PMC article.
-
RUNX2 promotes the suppression of osteoblast function and enhancement of osteoclast activity by multiple myeloma cells.Med Oncol. 2023 Mar 10;40(4):115. doi: 10.1007/s12032-023-01960-8. Med Oncol. 2023. PMID: 36897488 Free PMC article.
-
A New Model of Esophageal Cancers by Using a Detergent-Free Decellularized Matrix in a Perfusion Bioreactor.Bioengineering (Basel). 2023 Jan 11;10(1):96. doi: 10.3390/bioengineering10010096. Bioengineering (Basel). 2023. PMID: 36671668 Free PMC article.
References
-
- Adesunloye BA, Karzai FH, Dahut WL. Angiogenesis inhibitors in the treatment of prostate cancer. In: Marone G, Granata F, editors. Angiogenesis, Lymphangiogenesis and Clinical Implications. Basel: S. KARGER AG; 2013. pp. 197–215. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
