Mutations in Metabotropic Glutamate Receptor 1 Contribute to Natural Short Sleep Trait
- PMID: 33065013
- PMCID: PMC12352508
- DOI: 10.1016/j.cub.2020.09.071
Mutations in Metabotropic Glutamate Receptor 1 Contribute to Natural Short Sleep Trait
Abstract
Sufficient and efficient sleep is crucial for our health. Natural short sleepers can sleep significantly shorter than the average population without a desire for more sleep and without any obvious negative health consequences. In searching for genetic variants underlying the short sleep trait, we found two different mutations in the same gene (metabotropic glutamate receptor 1) from two independent natural short sleep families. In vitro, both of the mutations exhibited loss of function in receptor-mediated signaling. In vivo, the mice carrying the individual mutations both demonstrated short sleep behavior. In brain slices, both of the mutations changed the electrical properties and increased excitatory synaptic transmission. These results highlight the important role of metabotropic glutamate receptor 1 in modulating sleep duration.
Keywords: loss-of-function; mGluR1; short-sleep.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures
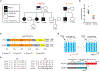



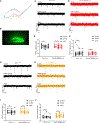

References
-
- Hirshkowitz M, Whiton K, Albert SM, Alessi C, Bruni O, DonCarlos L, Hazen N, Herman PJ, Adams Hillard PJ, Katz ES, et al. (2015). National Sleep Foundation’s updated sleep duration recommendations: final report. Sleep Health 1, 233–243. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

