RNA-Binding RING E3-Ligase DZIP3/hRUL138 Stabilizes Cyclin D1 to Drive Cell-Cycle and Cancer Progression
- PMID: 33067265
- PMCID: PMC7116596
- DOI: 10.1158/0008-5472.CAN-20-1871
RNA-Binding RING E3-Ligase DZIP3/hRUL138 Stabilizes Cyclin D1 to Drive Cell-Cycle and Cancer Progression
Abstract
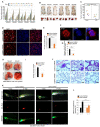
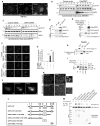
DZIP3/hRUL138 is a poorly characterized RNA-binding RING E3-ubiquitin ligase with functions in embryonic development. Here we demonstrate that DZIP3 is a crucial driver of cancer cell growth, migration, and invasion. In mice and zebrafish cancer models, DZIP3 promoted tumor growth and metastasis. In line with these results, DZIP3 was frequently overexpressed in several cancer types. Depletion of DZIP3 from cells resulted in reduced expression of Cyclin D1 and a subsequent G1 arrest and defect in cell growth. Mechanistically, DZIP3 utilized its two different domains to interact and stabilize Cyclin D1 both at mRNA and protein levels. Using an RNA-binding lysine-rich region, DZIP3 interacted with the AU-rich region in 3' untranslated region of Cyclin D1 mRNA and stabilized it. Using a RING E3-ligase domain, DZIP3 interacted and increased K63-linked ubiquitination of Cyclin D1 protein to stabilize it. Remarkably, DZIP3 interacted with, ubiquitinated, and stabilized Cyclin D1 predominantly in the G1 phase of the cell cycle, where it is needed for cell-cycle progression. In agreement with this, a strong positive correlation of mRNA expression between DZIP3 and Cyclin D1 in different cancer types was observed. Additionally, DZIP3 regulated several cell cycle proteins by modulating the Cyclin D1-E2F axes. Taken together, this study demonstrates for the first time that DZIP3 uses a unique two-pronged mechanism in its stabilization of Cyclin D1 to drive cell-cycle and cancer progression. SIGNIFICANCE: These findings show that DZIP3 is a novel driver of cell-cycle and cancer progression via its control of Cyclin D1 mRNA and protein stability in a cell-cycle phase-dependent manner. GRAPHICAL ABSTRACT: http://cancerres.aacrjournals.org/content/canres/81/2/315/F1.large.jpg.
©2020 American Association for Cancer Research.
Conflict of interest statement
The authors declare that they have no conflict of interests.
Figures





Similar articles
-
X-linked inhibitor of apoptosis protein (XIAP) regulation of cyclin D1 protein expression and cancer cell anchorage-independent growth via its E3 ligase-mediated protein phosphatase 2A/c-Jun axis.J Biol Chem. 2013 Jul 12;288(28):20238-47. doi: 10.1074/jbc.M112.448365. Epub 2013 May 29. J Biol Chem. 2013. PMID: 23720779 Free PMC article.
-
Circ-UBR1 facilitates proliferation, metastasis, and inhibits apoptosis in breast cancer by regulating the miR-1299/CCND1 axis.Life Sci. 2021 Feb 1;266:118829. doi: 10.1016/j.lfs.2020.118829. Epub 2020 Nov 28. Life Sci. 2021. PMID: 33259864
-
Roquin1 inhibits the proliferation of breast cancer cells by inducing G1/S cell cycle arrest via selectively destabilizing the mRNAs of cell cycle-promoting genes.J Exp Clin Cancer Res. 2020 Nov 23;39(1):255. doi: 10.1186/s13046-020-01766-w. J Exp Clin Cancer Res. 2020. PMID: 33228782 Free PMC article.
-
RN181 is a tumour suppressor in gastric cancer by regulation of the ERK/MAPK-cyclin D1/CDK4 pathway.J Pathol. 2019 Jun;248(2):204-216. doi: 10.1002/path.5246. Epub 2019 Apr 11. J Pathol. 2019. PMID: 30714150 Free PMC article.
-
IMP3 accelerates the progression of prostate cancer through inhibiting PTEN expression in a SMURF1-dependent way.J Exp Clin Cancer Res. 2020 Sep 16;39(1):190. doi: 10.1186/s13046-020-01657-0. J Exp Clin Cancer Res. 2020. Retraction in: J Exp Clin Cancer Res. 2023 Jan 17;42(1):24. doi: 10.1186/s13046-023-02599-z. PMID: 32938489 Free PMC article. Retracted.
Cited by
-
DZIP1 Expression as a Prognostic Marker in Gastric Cancer: A Bioinformatics-Based Analysis.Pharmgenomics Pers Med. 2021 Sep 16;14:1151-1168. doi: 10.2147/PGPM.S325701. eCollection 2021. Pharmgenomics Pers Med. 2021. PMID: 34557018 Free PMC article.
-
cIAP1-based degraders induce degradation via branched ubiquitin architectures.Nat Chem Biol. 2023 Mar;19(3):311-322. doi: 10.1038/s41589-022-01178-1. Epub 2022 Oct 31. Nat Chem Biol. 2023. PMID: 36316570
-
Emerging Roles of Non-proteolytic Ubiquitination in Tumorigenesis.Front Cell Dev Biol. 2022 Jul 6;10:944460. doi: 10.3389/fcell.2022.944460. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35874839 Free PMC article. Review.
-
Pan-cancer analysis uncovered the prognostic and therapeutic value of disulfidptosis.NPJ Precis Oncol. 2025 Feb 24;9(1):50. doi: 10.1038/s41698-025-00834-8. NPJ Precis Oncol. 2025. PMID: 39994355 Free PMC article.
-
Ring-finger protein RNF126 promotes prostate cancer progression via regulation of MBNL1.Sci Rep. 2025 Jul 4;15(1):23847. doi: 10.1038/s41598-025-04629-6. Sci Rep. 2025. PMID: 40615482 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

