Succession of Gut Microbial Structure in Twin Giant Pandas During the Dietary Change Stage and Its Role in Polysaccharide Metabolism
- PMID: 33072012
- PMCID: PMC7537565
- DOI: 10.3389/fmicb.2020.551038
Succession of Gut Microbial Structure in Twin Giant Pandas During the Dietary Change Stage and Its Role in Polysaccharide Metabolism
Abstract
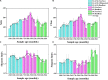
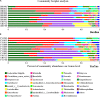
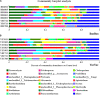
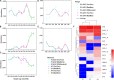
Adaptation to a bamboo diet is an essential process for giant panda growth, and gut microbes play an important role in the digestion of the polysaccharides in bamboo. The dietary transition in giant panda cubs is particularly complex, but it is an ideal period in which to study the effects of gut microbes on polysaccharide use because their main food changes from milk to bamboo (together with some bamboo shoot and coarse pastry). Here, we used 16S rDNA and internal transcribed spacer 1 (ITS1) DNA sequencing and metagenomic sequencing analysis to investigate the succession of the gut microbial structure in feces sampled from twin giant panda cubs during the completely dietary transition and determine the abundances of polysaccharide-metabolizing genes and their corresponding microbes to better understand the degradation of bamboo polysaccharides. Successive changes in the gut microbial diversity and structure were apparent in the growth of pandas during dietary shift process. Microbial diversity increased after the introduction of supplementary foods and then varied in a complex way for 1.5-2 years as bamboo and complex food components were introduced. They then stabilized after 2 years, when the cubs consumed a specialized bamboo diet. The microbes had more potential to metabolize the cellulose in bamboo than the hemicellulose, providing genes encoding cellulase systems corresponding to glycoside hydrolases (GHs; such as GH1, GH3, GH5, GH8, GH9, GH74, and GH94). The cellulose-metabolizing species (or genes) of gut bacteria was more abundant than that of gut fungi. Although cellulose-metabolizing species did not predominate in the gut bacterial community, microbial interactions allowed the giant pandas to achieve the necessary dietary shift and ultimately adapt to a bamboo diet.
Keywords: Clostridium; GH5; cellulose degradation; functional gene prediction; giant panda cub; gut microbes; succession.
Copyright © 2020 Zhan, Wang, Xie, Fu, Zhang, Wang, Zhou, Xu and Zhang.
Figures

Similar articles
-
An amateur gut microbial configuration formed in giant panda for striving to digest cellulose in bamboo: Systematic evidence from intestinal digestive enzymes, functional genes and microbial structures.Front Microbiol. 2022 Jul 26;13:926515. doi: 10.3389/fmicb.2022.926515. eCollection 2022. Front Microbiol. 2022. PMID: 35958139 Free PMC article.
-
Age-related alterations in metabolome and microbiome provide insights in dietary transition in giant pandas.mSystems. 2023 Jun 29;8(3):e0025223. doi: 10.1128/msystems.00252-23. Epub 2023 Jun 5. mSystems. 2023. PMID: 37273228 Free PMC article.
-
The carnivorous digestive system and bamboo diet of giant pandas may shape their low gut bacterial diversity.Conserv Physiol. 2020 Mar 13;8(1):coz104. doi: 10.1093/conphys/coz104. eCollection 2020. Conserv Physiol. 2020. PMID: 32190328 Free PMC article.
-
Progress in Research on the Gut Microflora of the Red Panda (Ailurus fulgens).Microorganisms. 2024 Feb 27;12(3):478. doi: 10.3390/microorganisms12030478. Microorganisms. 2024. PMID: 38543529 Free PMC article. Review.
-
Giant pandas are not an evolutionary cul-de-sac: evidence from multidisciplinary research.Mol Biol Evol. 2015 Jan;32(1):4-12. doi: 10.1093/molbev/msu278. Epub 2014 Oct 1. Mol Biol Evol. 2015. PMID: 25274274 Review.
Cited by
-
Composition, Influencing Factors, and Effects on Host Nutrient Metabolism of Fungi in Gastrointestinal Tract of Monogastric Animals.Animals (Basel). 2025 Mar 1;15(5):710. doi: 10.3390/ani15050710. Animals (Basel). 2025. PMID: 40075993 Free PMC article. Review.
-
An amateur gut microbial configuration formed in giant panda for striving to digest cellulose in bamboo: Systematic evidence from intestinal digestive enzymes, functional genes and microbial structures.Front Microbiol. 2022 Jul 26;13:926515. doi: 10.3389/fmicb.2022.926515. eCollection 2022. Front Microbiol. 2022. PMID: 35958139 Free PMC article.
-
Succession of Intestinal Microbial Structure of Giant Pandas (Ailuropoda melanoleuca) during Different Developmental Stages and Its Correlation with Cellulase Activity.Animals (Basel). 2021 Aug 10;11(8):2358. doi: 10.3390/ani11082358. Animals (Basel). 2021. PMID: 34438815 Free PMC article.
-
Integrated multi-omics reveals novel microbe-host lipid metabolism and immune interactions in the donkey hindgut.Front Immunol. 2022 Nov 18;13:1003247. doi: 10.3389/fimmu.2022.1003247. eCollection 2022. Front Immunol. 2022. PMID: 36466834 Free PMC article.
-
Age-related alterations in metabolome and microbiome provide insights in dietary transition in giant pandas.mSystems. 2023 Jun 29;8(3):e0025223. doi: 10.1128/msystems.00252-23. Epub 2023 Jun 5. mSystems. 2023. PMID: 37273228 Free PMC article.
References
-
- Anand A. A., Chattopadhyay B., Kandula S. (2012). Isolation and characterization of cellulose-degrading and xylanolytic bacteria from the short-nosed fruit bat Cynopterus sphinx. Acta Chiropterol. 14 233–239.
-
- Bhat I. U., Mustafa M. T., Mohmod A. L., Abdul Khalil H. P. S. (2011). Specteroscopic, thermal, and anatomical characterization of cultivated of cultivated bamboo (Gigantochloa spp.). Bioresources 6 1752–1763.
LinkOut - more resources
Full Text Sources

