Age-associated mitochondrial DNA mutations cause metabolic remodelling that contributes to accelerated intestinal tumorigenesis
- PMID: 33073241
- PMCID: PMC7116185
- DOI: 10.1038/s43018-020-00112-5
Age-associated mitochondrial DNA mutations cause metabolic remodelling that contributes to accelerated intestinal tumorigenesis
Erratum in
-
Author Correction: Age-associated mitochondrial DNA mutations cause metabolic remodeling that contributes to accelerated intestinal tumorigenesis.Nat Cancer. 2021 Jan;2(1):129. doi: 10.1038/s43018-020-00156-7. Nat Cancer. 2021. PMID: 35121898 No abstract available.
Abstract
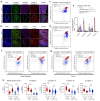
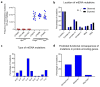
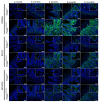
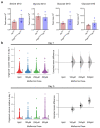
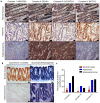
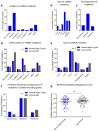
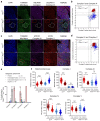
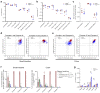
Oxidative phosphorylation (OXPHOS) defects caused by somatic mitochondrial DNA (mtDNA) mutations increase with age in human colorectal epithelium and are prevalent in colorectal tumours, but whether they actively contribute to tumorigenesis remains unknown. Here we demonstrate that mtDNA mutations causing OXPHOS defects are enriched during the human adenoma/carcinoma sequence, suggesting they may confer a metabolic advantage. To test this we deleted the tumour suppressor Apc in OXPHOS deficient intestinal stem cells in mice. The resulting tumours were larger than in control mice due to accelerated cell proliferation and reduced apoptosis. We show that both normal crypts and tumours undergo metabolic remodelling in response to OXPHOS deficiency by upregulating the de novo serine synthesis pathway (SSP). Moreover, normal human colonic crypts upregulate the SSP in response to OXPHOS deficiency prior to tumorigenesis. Our data show that age-associated OXPHOS deficiency causes metabolic remodelling that can functionally contribute to accelerated intestinal cancer development.
Conflict of interest statement
Competing interests: F.O is a director of Fibrofind limited. J.L and F.O are shareholders in Fibrofind limited. The remaining authors declare no competing financial interests.
Figures





Comment in
-
Metabolic benefits of mitochondrial DNA mutations.Nat Rev Cancer. 2020 Dec;20(12):696. doi: 10.1038/s41568-020-00309-x. Nat Rev Cancer. 2020. PMID: 33024262 No abstract available.
-
MtDNA mutations and aging-not a closed case after all?Signal Transduct Target Ther. 2021 Feb 10;6(1):56. doi: 10.1038/s41392-021-00479-6. Signal Transduct Target Ther. 2021. PMID: 33563891 Free PMC article. No abstract available.
-
mtDNA mutations help support cancer cells.Nat Cancer. 2020 Oct;1(10):941-942. doi: 10.1038/s43018-020-00128-x. Nat Cancer. 2020. PMID: 35121873 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
- MR/L016354/1/MRC_/Medical Research Council/United Kingdom
- BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 21139/CRUK_/Cancer Research UK/United Kingdom
- MR/T015586/1/MRC_/Medical Research Council/United Kingdom
- MR/K001949/1/MRC_/Medical Research Council/United Kingdom
- A12481/CRUK_/Cancer Research UK/United Kingdom
- A23390/CRUK_/Cancer Research UK/United Kingdom
- MR/R023026/1/MRC_/Medical Research Council/United Kingdom
- A25045/CRUK_/Cancer Research UK/United Kingdom
- 203105/WT_/Wellcome Trust/United Kingdom
- 311301/ERC_/European Research Council/International
- A21139/CRUK_/Cancer Research UK/United Kingdom
- C18342/A23390/CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Medical

