A high-quality Brassica napus genome reveals expansion of transposable elements, subgenome evolution and disease resistance
- PMID: 33073445
- PMCID: PMC7955885
- DOI: 10.1111/pbi.13493
A high-quality Brassica napus genome reveals expansion of transposable elements, subgenome evolution and disease resistance
Abstract
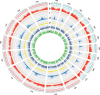
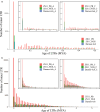
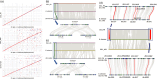
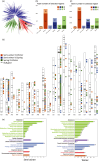
Rapeseed (Brassica napus L.) is a recent allotetraploid crop, which is well known for its high oil production. Here, we report a high-quality genome assembly of a typical semi-winter rapeseed cultivar, 'Zhongshuang11' (hereafter 'ZS11'), using a combination of single-molecule sequencing and chromosome conformation capture (Hi-C) techniques. Most of the high-confidence sequences (93.1%) were anchored to the individual chromosomes with a total of 19 centromeres identified, matching the exact chromosome count of B. napus. The repeat sequences in the A and C subgenomes in B. napus expanded significantly from 500 000 years ago, especially over the last 100 000 years. These young and recently amplified LTR-RTs showed dispersed chromosomal distribution but significantly preferentially clustered into centromeric regions. We exhaustively annotated the nucleotide-binding leucine-rich repeat (NLR) gene repertoire, yielding a total of 597 NLR genes in B. napus genome and 17.4% of which are paired (head-to-head arrangement). Based on the resequencing data of 991 B. napus accessions, we have identified 18 759 245 single nucleotide polymorphisms (SNPs) and detected a large number of genomic regions under selective sweep among the three major ecotype groups (winter, semi-winter and spring) in B. napus. We found 49 NLR genes and five NLR gene pairs colocated in selective sweep regions with different ecotypes, suggesting a rapid diversification of NLR genes during the domestication of B. napus. The high quality of our B. napus 'ZS11' genome assembly could serve as an important resource for the study of rapeseed genomics and reveal the genetic variations associated with important agronomic traits.
Keywords: Brassica napus; disease resistance; long-read sequencing; selective sweep; subgenome evolution.
© 2020 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declared that they have no conflict of interest to this work.
Figures


References
-
- Abrusan, G. , Grundmann, N. , DeMester, L. and Makalowski, W. (2009) TEclass–a tool for automated classification of unknown eukaryotic transposable elements. Bioinformatics, 25, 1329–1330. - PubMed
-
- Bayer, P.E. , Edwards, D. and Batley, J. (2018) Bias in resistance gene prediction due to repeat masking. Nat Plants, 4, 762–765. - PubMed
-
- Bennetzen, J.L. and Wang, H. (2014) The contributions of transposable elements to the structure, function, and evolution of plant genomes. Annu. Rev. Plant Biol. 65, 505–530. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

