Genome sequencing increases diagnostic yield in clinically diagnosed Alagille syndrome patients with previously negative test results
- PMID: 33077891
- PMCID: PMC7862053
- DOI: 10.1038/s41436-020-00989-8
Genome sequencing increases diagnostic yield in clinically diagnosed Alagille syndrome patients with previously negative test results
Abstract
Purpose: Detection of all major classes of genomic variants in a single test would decrease cost and increase the efficiency of genomic diagnostics. Genome sequencing (GS) has the potential to provide this level of comprehensive detection. We sought to demonstrate the utility of GS in the molecular diagnosis of 18 patients with clinically defined Alagille syndrome (ALGS), who had a negative or inconclusive result by standard-of-care testing.
Methods: We performed GS on 16 pathogenic variant-negative probands and two probands with inconclusive results (of 406 ALGS probands) and analyzed the data for sequence, copy-number, and structural variants in JAG1 and NOTCH2.
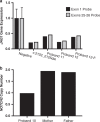
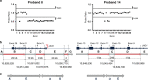
Results: GS identified four novel pathogenic alterations including a copy-neutral inversion, a partial deletion, and a promoter variant in JAG1, and a partial NOTCH2 deletion, for an additional diagnostic yield of 0.9%. Furthermore, GS resolved two complex rearrangements, resulting in identification of a pathogenic variant in 97.5% (n = 396/406) of patients after GS.
Conclusion: GS provided an increased diagnostic yield for individuals with clinically defined ALGS who had prior negative or incomplete genetic testing by other methods. Our results show that GS can detect all major classes of variants and has potential to become a single first-tier diagnostic test for Mendelian disorders.
Keywords: Alagille syndrome; JAG1; NOTCH2, diagnostic testing; genome sequencing.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Comment in
-
Functional characterization of a JAG1 5'UTR variant in a patient with clinically observed Alagille syndrome.Clin Genet. 2022 Sep;102(3):248-250. doi: 10.1111/cge.14179. Epub 2022 Jun 27. Clin Genet. 2022. PMID: 35761784 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

