The tree of life of polyamine oxidases
- PMID: 33082384
- PMCID: PMC7576179
- DOI: 10.1038/s41598-020-74708-3
The tree of life of polyamine oxidases
Abstract
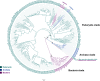
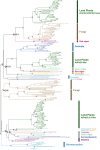
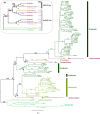
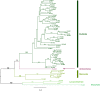
Polyamine oxidases (PAOs) are characterized by a broad variability in catalytic properties and subcellular localization, and impact key cellular processes in diverse organisms. In the present study, a comprehensive phylogenetic analysis was performed to understand the evolution of PAOs across the three domains of life and particularly within eukaryotes. Phylogenetic trees show that PAO-like sequences of bacteria, archaea, and eukaryotes form three distinct clades, with the exception of a few procaryotes that probably acquired a PAO gene through horizontal transfer from a eukaryotic donor. Results strongly support a common origin for archaeal PAO-like proteins and eukaryotic PAOs, as well as a shared origin between PAOs and monoamine oxidases. Within eukaryotes, four main lineages were identified that likely originated from an ancestral eukaryotic PAO before the split of the main superphyla, followed by specific gene losses in each superphylum. Plant PAOs show the highest diversity within eukaryotes and belong to three distinct clades that underwent to multiple events of gene duplication and gene loss. Peptide deletion along the evolution of plant PAOs of Clade I accounted for further diversification of function and subcellular localization. This study provides a reference for future structure-function studies and emphasizes the importance of extending comparisons among PAO subfamilies across multiple eukaryotic superphyla.
Conflict of interest statement
The authors declare no competing interests.
Figures
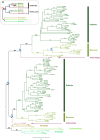
References
-
- Hamana K, Niitsu M, Samejima K, Matsuzaki S. Linear and branched pentaamines, hexaamines and heptaamines in seeds of Vicia sativa. Phytochemistry. 1991;30:3319–3322. doi: 10.1016/0031-9422(91)83201-U. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

