Impact of mixed biofilm formation with environmental microorganisms on E. coli O157:H7 survival against sanitization
- PMID: 33083548
- PMCID: PMC7560865
- DOI: 10.1038/s41538-020-00076-x
Impact of mixed biofilm formation with environmental microorganisms on E. coli O157:H7 survival against sanitization
Abstract
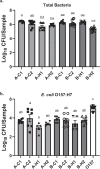
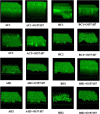
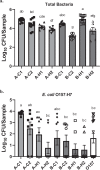
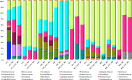
Biofilm formation by foodborne pathogens is a serious threat to food safety and public health. Meat processing plants may harbor various microorganisms and occasional foodborne pathogens; thus, the environmental microbial community might impact pathogen survival via mixed biofilm formation. We collected floor drain samples from two beef plants with different E. coli O157:H7 prevalence history and investigated the effects of the environmental microorganisms on pathogen sanitizer tolerance. The results showed that biofilm forming ability and bacterial species composition varied considerably based on the plants and drain locations. E. coli O157:H7 cells obtained significantly higher sanitizer tolerance in mixed biofilms by samples from the plant with recurrent E. coli O157:H7 prevalence than those mixed with samples from the other plant. The mixed biofilm that best protected E. coli O157:H7 also had the highest species diversity. The percentages of the species were altered significantly after sanitization, suggesting that the community composition affects the role and tolerance level of each individual species. Therefore, the unique environmental microbial community, their ability to form biofilms on contact surfaces and the interspecies interactions all play roles in E. coli O157:H7 persistence by either enhancing or reducing pathogen survival within the biofilm community.
Keywords: Biofilms; Industrial microbiology.
© This is a U.S. government work and not under copyright protection in the U.S.; foreign copyright protection may apply 2020.
Conflict of interest statement
Competing interestsThe authors declare no competing interests.
Figures
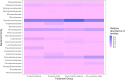
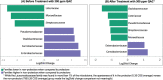
References
-
- Wang R, Kalchayanand N, Schmidt JW, Harhay DM. Mixed biofilm formation by Shiga toxin-producing Escherichia coli and Salmonella enterica serovar Typhimurium enhanced bacterial resistance to sanitization due to extracellular polymeric substances. J. Food Prot. 2013;76:1513–1522. doi: 10.4315/0362-028X.JFP-13-077. - DOI - PubMed
-
- Chitlapilly Dass S, Abu-Ghannam N, Antony-Babu S, Cummins JE. Ecology and molecular typing of L. monocytogenes in a processing plant for cold-smoked salmon in the Republic of Ireland. Food Res. Int. 2010;43:1529–1536. doi: 10.1016/j.foodres.2010.04.030. - DOI
LinkOut - more resources
Full Text Sources

