Overexpression of CDSP32 (GhTRX134) Cotton Gene Enhances Drought, Salt, and Oxidative Stress Tolerance in Arabidopsis
- PMID: 33086523
- PMCID: PMC7650641
- DOI: 10.3390/plants9101388
Overexpression of CDSP32 (GhTRX134) Cotton Gene Enhances Drought, Salt, and Oxidative Stress Tolerance in Arabidopsis
Abstract
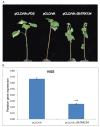
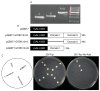
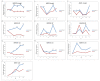
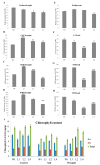
Upland cotton (Gossypium hirsutum L.) is the main natural fiber crop worldwide and is an essential source of seed oil and biofuel products. Many abiotic stresses, such as drought and salinity, constrain cotton production. Thioredoxins (TRXs) are a group of small ubiquitous proteins that are widely distributed among organisms. TRXs play a crucial role in regulating diverse functions during plant growth and development. In the present study, a novel GhTRX134 gene was characterized and overexpressed in Arabidopsis and silenced in cotton under drought stress. Furthermore, the proline content and enzyme activity levels were measured in transgenic plants and wild-type (Wt) plants under drought and salt stress. The results revealed that the overexpression of GhTRX134 enhanced abiotic stress tolerance. When GhTRX134 was silenced, cotton plants become more sensitive to drought. Taken together, these findings confirmed that the overexpression of GhTRX134 improved drought and salt tolerance in Arabidopsis plants. Therefore, the GhTRX134 gene can be transformed into cotton plants to obtain transgenic lines for more functional details.
Keywords: Arabidopsis; abiotic stress; gene cloning; thioredoxins; transformation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Overexpression of StDREB2 Transcription Factor Enhances Drought Stress Tolerance in Cotton (Gossypium barbadense L.).Genes (Basel). 2019 Feb 14;10(2):142. doi: 10.3390/genes10020142. Genes (Basel). 2019. PMID: 30769841 Free PMC article.
-
Cotton GhERF38 gene is involved in plant response to salt/drought and ABA.Ecotoxicology. 2017 Aug;26(6):841-854. doi: 10.1007/s10646-017-1815-2. Epub 2017 May 23. Ecotoxicology. 2017. PMID: 28536792
-
The cotton WRKY transcription factor GhWRKY17 functions in drought and salt stress in transgenic Nicotiana benthamiana through ABA signaling and the modulation of reactive oxygen species production.Plant Cell Physiol. 2014 Dec;55(12):2060-76. doi: 10.1093/pcp/pcu133. Epub 2014 Sep 26. Plant Cell Physiol. 2014. PMID: 25261532
-
Regulatory Network of Cotton Genes in Response to Salt, Drought and Wilt Diseases (Verticillium and Fusarium): Progress and Perspective.Front Plant Sci. 2021 Nov 29;12:759245. doi: 10.3389/fpls.2021.759245. eCollection 2021. Front Plant Sci. 2021. PMID: 34912357 Free PMC article. Review.
-
Mechanism of cotton resistance to abiotic stress, and recent research advances in the osmoregulation related genes.Front Plant Sci. 2022 Aug 17;13:972635. doi: 10.3389/fpls.2022.972635. eCollection 2022. Front Plant Sci. 2022. PMID: 36061778 Free PMC article. Review.
Cited by
-
Functional Characterization of Lobularia maritima LmTrxh2 Gene Involved in Cold Tolerance in Tobacco through Alleviation of ROS Damage to the Plasma Membrane.Int J Mol Sci. 2023 Feb 3;24(3):3030. doi: 10.3390/ijms24033030. Int J Mol Sci. 2023. PMID: 36769352 Free PMC article.
-
Deciphering the complex cotton genome for improving fiber traits and abiotic stress resilience in sustainable agriculture.Mol Biol Rep. 2023 Aug;50(8):6937-6953. doi: 10.1007/s11033-023-08565-4. Epub 2023 Jun 22. Mol Biol Rep. 2023. PMID: 37349608 Review.
-
Integrating genes and metabolites: unraveling mango's drought resilience mechanisms.BMC Plant Biol. 2024 Mar 23;24(1):208. doi: 10.1186/s12870-024-04908-w. BMC Plant Biol. 2024. PMID: 38519933 Free PMC article.
-
Overexpression of cotton GhNAC072 gene enhances drought and salt stress tolerance in transgenic Arabidopsis.BMC Genomics. 2022 Sep 12;23(1):648. doi: 10.1186/s12864-022-08876-z. BMC Genomics. 2022. PMID: 36096725 Free PMC article.
-
Physiological response and drought resistance evaluation of Gleditsia sinensis seedlings under drought-rehydration state.Sci Rep. 2023 Nov 15;13(1):19963. doi: 10.1038/s41598-023-45394-8. Sci Rep. 2023. PMID: 37968307 Free PMC article.
References
-
- Wang X., Yu Y., Sang J., Wu Q., Zhang X., Lin Z. Intraspecific linkage map construction and QTL mapping of yield and fiber quality of ‘Gossypiumbabrdense’. [(accessed on 3 March 2020)];Aust. J. Crop. Sci. 2013 7:1252–1261. Available online: http://search.informit.com.au/documentSummary;dn=625250539555035;res=IELHSS.
-
- Elasad M., Wei H., Wang H., Su J., Ondati E., Yu S. Genome-Wide Analysis and Characterization of the TRX Gene Family in Upland Cotton. Trop. Plant Biol. 2018;11:119–130. doi: 10.1007/s12042-018-9205-3. - DOI
-
- Zhao G., Song Y., Wang C., Butt H.I., Wang Q., Zhang C., Yang Z., Liu Z., Chen E., Zhang X., et al. Genome-wide identification and functional analysis of the TIFY gene family in response to drought in cotton. Mol. Genet. Genom. 2016;291:2173–2187. doi: 10.1007/s00438-016-1248-2. - DOI - PMC - PubMed
-
- Colville L., Kranner I. Desiccation tolerant plants as model systems to study redox regulation of protein thiols. Plant. Growth Regul. 2010;62:241–255. doi: 10.1007/s10725-010-9482-9. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

