Synthetic gene-regulatory networks in the opportunistic human pathogen Streptococcus pneumoniae
- PMID: 33087560
- PMCID: PMC7959565
- DOI: 10.1073/pnas.1920015117
Synthetic gene-regulatory networks in the opportunistic human pathogen Streptococcus pneumoniae
Abstract
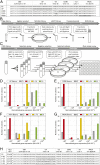
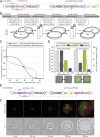
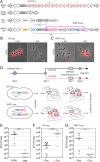
Streptococcus pneumoniae can cause disease in various human tissues and organs, including the ear, the brain, the blood, and the lung, and thus in highly diverse and dynamic environments. It is challenging to study how pneumococci control virulence factor expression, because cues of natural environments and the presence of an immune system are difficult to simulate in vitro. Here, we apply synthetic biology methods to reverse-engineer gene expression control in S. pneumoniae A selection platform is described that allows for straightforward identification of transcriptional regulatory elements out of combinatorial libraries. We present TetR- and LacI-regulated promoters that show expression ranges of four orders of magnitude. Based on these promoters, regulatory networks of higher complexity are assembled, such as logic AND gates and IMPLY gates. We demonstrate single-copy genome-integrated toggle switches that give rise to bimodal population distributions. The tools described here can be used to mimic complex expression patterns, such as the ones found for pneumococcal virulence factors. Indeed, we were able to rewire gene expression of the capsule operon, the main pneumococcal virulence factor, to be externally inducible (YES gate) or to act as an IMPLY gate (only expressed in absence of inducer). Importantly, we demonstrate that these synthetic gene-regulatory networks are functional in an influenza A virus superinfection murine model of pneumonia, paving the way for in vivo investigations of the importance of gene expression control on the pathogenicity of S. pneumoniae.
Keywords: Pneumococcus; counterselection; superinfection; synthetic biology; toggle switch.
Copyright © 2020 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures



Similar articles
-
Exploration of Bacterial Bottlenecks and Streptococcus pneumoniae Pathogenesis by CRISPRi-Seq.Cell Host Microbe. 2021 Jan 13;29(1):107-120.e6. doi: 10.1016/j.chom.2020.10.001. Epub 2020 Oct 28. Cell Host Microbe. 2021. PMID: 33120116 Free PMC article.
-
The core promoter of the capsule operon of Streptococcus pneumoniae is necessary for colonization and invasive disease.Infect Immun. 2014 Feb;82(2):694-705. doi: 10.1128/IAI.01289-13. Epub 2013 Nov 25. Infect Immun. 2014. PMID: 24478084 Free PMC article.
-
Gene expression platform for synthetic biology in the human pathogen Streptococcus pneumoniae.ACS Synth Biol. 2015 Mar 20;4(3):228-39. doi: 10.1021/sb500229s. Epub 2014 Jun 4. ACS Synth Biol. 2015. PMID: 24845455
-
Virulence factors in pneumococcal respiratory pathogenesis.Future Microbiol. 2008 Apr;3(2):205-21. doi: 10.2217/17460913.3.2.205. Future Microbiol. 2008. PMID: 18366340 Review.
-
Streptococcus pneumoniae two-component regulatory systems: The interplay of the pneumococcus with its environment.Int J Med Microbiol. 2018 Aug;308(6):722-737. doi: 10.1016/j.ijmm.2017.11.012. Epub 2017 Nov 26. Int J Med Microbiol. 2018. PMID: 29221986 Review.
Cited by
-
Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions.Nat Commun. 2023 Nov 17;14(1):7454. doi: 10.1038/s41467-023-43241-y. Nat Commun. 2023. PMID: 37978173 Free PMC article.
-
Make-or-break prime editing for genome engineering in Streptococcus pneumoniae.Nat Commun. 2025 Apr 23;16(1):3796. doi: 10.1038/s41467-025-59068-8. Nat Commun. 2025. PMID: 40263274 Free PMC article.
-
Competence remodels the pneumococcal cell wall exposing key surface virulence factors that mediate increased host adherence.PLoS Biol. 2023 Jan 30;21(1):e3001990. doi: 10.1371/journal.pbio.3001990. eCollection 2023 Jan. PLoS Biol. 2023. PMID: 36716340 Free PMC article.
-
CRISPRi-TnSeq maps genome-wide interactions between essential and non-essential genes in bacteria.Nat Microbiol. 2024 Sep;9(9):2395-2409. doi: 10.1038/s41564-024-01759-x. Epub 2024 Jul 19. Nat Microbiol. 2024. PMID: 39030344 Free PMC article.
-
Status quo of tet regulation in bacteria.Microb Biotechnol. 2022 Apr;15(4):1101-1119. doi: 10.1111/1751-7915.13926. Epub 2021 Oct 29. Microb Biotechnol. 2022. PMID: 34713957 Free PMC article. Review.
References
-
- Kadioglu A., Weiser J. N., Paton J. C., Andrew P. W., The role of Streptococcus pneumoniae virulence factors in host respiratory colonization and disease. Nat. Rev. Microbiol. 6, 288–301 (2008). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

