Exome sequencing and characterization of 49,960 individuals in the UK Biobank
- PMID: 33087929
- PMCID: PMC7759458
- DOI: 10.1038/s41586-020-2853-0
Exome sequencing and characterization of 49,960 individuals in the UK Biobank
Abstract
The UK Biobank is a prospective study of 502,543 individuals, combining extensive phenotypic and genotypic data with streamlined access for researchers around the world1. Here we describe the release of exome-sequence data for the first 49,960 study participants, revealing approximately 4 million coding variants (of which around 98.6% have a frequency of less than 1%). The data include 198,269 autosomal predicted loss-of-function (LOF) variants, a more than 14-fold increase compared to the imputed sequence. Nearly all genes (more than 97%) had at least one carrier with a LOF variant, and most genes (more than 69%) had at least ten carriers with a LOF variant. We illustrate the power of characterizing LOF variants in this population through association analyses across 1,730 phenotypes. In addition to replicating established associations, we found novel LOF variants with large effects on disease traits, including PIEZO1 on varicose veins, COL6A1 on corneal resistance, MEPE on bone density, and IQGAP2 and GMPR on blood cell traits. We further demonstrate the value of exome sequencing by surveying the prevalence of pathogenic variants of clinical importance, and show that 2% of this population has a medically actionable variant. Furthermore, we characterize the penetrance of cancer in carriers of pathogenic BRCA1 and BRCA2 variants. Exome sequences from the first 49,960 participants highlight the promise of genome sequencing in large population-based studies and are now accessible to the scientific community.
Conflict of interest statement
C.V.V.H., J.D.B., D.L., C.G.-J., S.K., B.Y., N.B., A.H.L., C.O., A.M., J.S., C.S., A.H., E.M., L.B., A.L., X.B., S.O., J.P., L.H., A.L.B., A.Y., K.P., M.J., W.J.S., G.D.Y., A.E., G.C., A.R.S., S.B., M.C., J.G.R., J.M., J.D.O., G.R.A., A.B. and the spouse of C.J.W. are current or former employees and/or stockholders of Regeneron Genetics Center or Regeneron Pharmaceuticals. J.P. is a current employee of DNANexus and C.S. of Hasso Plattner Institute, but work was conducted while employed by the Regeneron Genetics Center. I.T., J.D.H., A.K.P., L.C., M.R.N., J.W., R.A.S. and L.Y.-A. are current or former employees and/or stockholders of GlaxoSmithKline. I.T. is a current employee of AstraZenica, J.D.H. of Foresite Labs, L.C. of BioMarin and M.R.N. of Deerfield, but work was conducted while employed by GlaxoSmithKline. The other authors declare no competing interests.
Figures
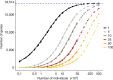
Comment in
-
Whole exome sequencing of large populations: identification of loss of function alleles and implications for inherited kidney diseases.Kidney Int. 2021 Jun;99(6):1255-1259. doi: 10.1016/j.kint.2020.12.036. Epub 2021 Feb 5. Kidney Int. 2021. PMID: 33549588 No abstract available.
-
How the human genome transformed study of rare diseases.Nature. 2021 Feb;590(7845):218-219. doi: 10.1038/d41586-021-00294-7. Nature. 2021. PMID: 33568830 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

