Association of molecular markers with physio-biochemical traits related to seed vigour in rice
- PMID: 33088044
- PMCID: PMC7548267
- DOI: 10.1007/s12298-020-00879-y
Association of molecular markers with physio-biochemical traits related to seed vigour in rice
Abstract
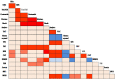
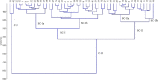
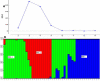
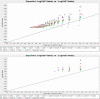
Eighteen physio-biochemical traits influencing seed vigour were studied for their association with molecular markers using a mini core set constituted from 120 germplasm lines. High genetic variation was detected in the parameters namely chlrophyll a, Chlrophyll b, total chlorophyll, carotenoids, total anthocyanin content, gamma-oryzanols, total phenolics content, superoxide dismutase, catalase, guaicol peroxidase, total soluble sugar, total protein, seed vigour index -I and seed vigour index -II. Strong positive correlation of seed vigour index II was observed with amylose content, total anthocyanin content, catalase, total phenolic content and total flavonoid content while a negative association was observed for gamma-oryzanol content. High gene diversity (0.7169) and informative markers value (0.6789) were estimated from the investigation. Three genetic structure groups were observed in the panel population and genotypes were grouped in the subpopulations based on the seed vigour trait. Differences in the fixation indices of the three sub populations indicated existence of linkage disequilibrium in the studied panel population. Association of the traits namely total flavonoids, superoxide dismutase, catalase, chlorophyll a, Chlorophyll b, total chlorophyll, carotenoids, starch, amylose, total anthocyanin, gamma-oryzanol, total phenolics with the molecular markers were detected by Generalized Linear Model and Mixed Linear Model showing > 0.10 R2 value. Association of the trait, total flavonoids with marker RM7364 located on chromosome 8 reported in earlier study was validated in this investigation. The validated markers and the novel markers detected showing higher R2 value will be useful for improvement of seed vigour in rice.
Keywords: Biochemical traits; Marker-trait association; Molecular markers; Physiological traits; Rice; Seed vigour.
© Prof. H.S. Srivastava Foundation for Science and Society 2020.
Conflict of interest statement
Competing interestsWe declare that there is no competing interest among the authors.
Figures
Similar articles
-
Association mapping for protein, total soluble sugars, starch, amylose and chlorophyll content in rice.BMC Plant Biol. 2022 Dec 29;22(1):620. doi: 10.1186/s12870-022-04015-8. BMC Plant Biol. 2022. PMID: 36581797 Free PMC article.
-
Unraveling the genomic regions controlling the seed vigour index, root growth parameters and germination per cent in rice.PLoS One. 2022 Jul 26;17(7):e0267303. doi: 10.1371/journal.pone.0267303. eCollection 2022. PLoS One. 2022. PMID: 35881571 Free PMC article.
-
Association mapping of grain color, phenolic content, flavonoid content and antioxidant capacity in dehulled rice.Theor Appl Genet. 2011 Mar;122(5):1005-16. doi: 10.1007/s00122-010-1505-4. Epub 2010 Dec 15. Theor Appl Genet. 2011. PMID: 21161500
-
Early seedling vigour, an imperative trait for direct-seeded rice: an overview on physio-morphological parameters and molecular markers.Planta. 2015 May;241(5):1027-50. doi: 10.1007/s00425-015-2273-9. Epub 2015 Mar 25. Planta. 2015. PMID: 25805338 Review.
-
Phenomics of rice early vigour and drought response: Are sugar related and morphogenetic traits relevant?Rice (N Y). 2012 Aug 20;5(1):22. doi: 10.1186/1939-8433-5-22. Rice (N Y). 2012. PMID: 24279832 Free PMC article.
Cited by
-
Association mapping for protein, total soluble sugars, starch, amylose and chlorophyll content in rice.BMC Plant Biol. 2022 Dec 29;22(1):620. doi: 10.1186/s12870-022-04015-8. BMC Plant Biol. 2022. PMID: 36581797 Free PMC article.
-
The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize.Int J Mol Sci. 2022 Nov 28;23(23):14865. doi: 10.3390/ijms232314865. Int J Mol Sci. 2022. PMID: 36499196 Free PMC article.
-
Integrated multispectral imaging, germination phenotype, and transcriptomic analysis provide insights into seed vigor responsive mechanisms in quinoa under artificial accelerated aging.Front Plant Sci. 2024 Sep 30;15:1435154. doi: 10.3389/fpls.2024.1435154. eCollection 2024. Front Plant Sci. 2024. PMID: 39403620 Free PMC article.
-
The Potential of High-Anthocyanin Purple Rice as a Functional Ingredient in Human Health.Antioxidants (Basel). 2021 May 24;10(6):833. doi: 10.3390/antiox10060833. Antioxidants (Basel). 2021. PMID: 34073767 Free PMC article. Review.
-
Population genetic structure and association mapping for iron toxicity tolerance in rice.PLoS One. 2021 Mar 1;16(3):e0246232. doi: 10.1371/journal.pone.0246232. eCollection 2021. PLoS One. 2021. PMID: 33647046 Free PMC article.
References
-
- Aebi H. Catalase in vitro methods. In: Colowick SP, Kaplan NO, editors. Methods in enzymology. Cambridge: Academic Press; 1984. p. 105, 114–121. - PubMed
-
- Agrama HA, Eizenga GC, Yan W. Association mapping of yield and its components in rice cultivars. Mol Breed. 2007;19:341–356. doi: 10.1007/s11032-006-9066-6. - DOI
-
- Allard RW. Principles of plant breeding. New York: Wiley; 1960.
-
- Anandan A, Pradhan SK, Das SK, Behera L, Sangeetha G. Differential responses of rice genotypes and physiological mechanism under prolonged deepwater flooding. Field Crops Res. 2015;172:153–163.
LinkOut - more resources
Full Text Sources
