Development of a novel secondary phenotypic screen to identify hits within the mycobacterial protein synthesis pipeline
- PMID: 33089076
- PMCID: PMC7566049
- DOI: 10.1096/fba.2020-00022
Development of a novel secondary phenotypic screen to identify hits within the mycobacterial protein synthesis pipeline
Abstract
Background: Whole-cell phenotypic screening is the driving force behind modern anti-tubercular drug discovery efforts. Focus has shifted from screening for bactericidal scaffolds to screens incorporating target deconvolution. Target-based screening aims to direct drug discovery toward known effective targets and avoid investing resources into unproductive lines of enquiry. The protein synthesis pipeline, including RNA polymerase and the ribosome, is a clinically proven target in Mycobacterium tuberculosis. Screening for new hits of this effective target pathway is an invaluable tool in the drug discovery arsenal.
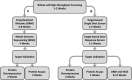
Methods: Using M. tuberculosis H37Rv augmented with anhydrotetracycline-inducible expression of mCherry, a phenotypic screen was developed for the identification of protein synthesis inhibitors in a medium throughput screening format.
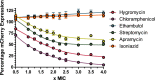
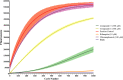
Results: The assay was validated using known inhibitors of protein synthesis to show a dose-dependent reduction in mCherry fluorescence. This was expanded to a proprietary screen of hypothetical protein synthesis hits and modified to include quantitative viability measurement of cells using resazurin.
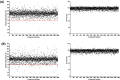
Conclusion: Following the success of the proprietary screen, a larger scale screen of the GlaxoSmithKline anti-tubercular library containing 2799 compounds was conducted. Combined single shot and dose-response screening yielded 18 hits, 0.64% of all screened compounds.
Keywords: RNA polymerase; mCherry; mycobacteria; ribosome; transcription; translation.
©2020 The Authors. FASEB BioAdvances published by The Federation of American Societies for Experimental Biology.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



Similar articles
-
Development of a whole-cell high-throughput phenotypic screen to identify inhibitors of mycobacterial amino acid biosynthesis.FASEB Bioadv. 2019 Jan 22;1(4):246-254. doi: 10.1096/fba.2018-00048. eCollection 2019 Apr. FASEB Bioadv. 2019. PMID: 32123830 Free PMC article.
-
Fragment-Based Whole Cell Screen Delivers Hits against M. tuberculosis and Non-tuberculous Mycobacteria.Front Microbiol. 2016 Sep 7;7:1392. doi: 10.3389/fmicb.2016.01392. eCollection 2016. Front Microbiol. 2016. PMID: 27656168 Free PMC article.
-
Development of Mycobacterium tuberculosis whole cell screening hits as potential antituberculosis agents.J Med Chem. 2013 Oct 24;56(20):7755-60. doi: 10.1021/jm400381v. Epub 2013 Sep 23. J Med Chem. 2013. PMID: 23927683 Review.
-
A high-throughput target-based screening approach for the identification and assessment of Mycobacterium tuberculosis mycothione reductase inhibitors.Microbiol Spectr. 2024 Mar 5;12(3):e0372323. doi: 10.1128/spectrum.03723-23. Epub 2024 Feb 5. Microbiol Spectr. 2024. PMID: 38315026 Free PMC article.
-
Perspective: Challenges and opportunities in TB drug discovery from phenotypic screening.Bioorg Med Chem. 2015 Aug 15;23(16):5087-97. doi: 10.1016/j.bmc.2014.12.031. Epub 2014 Dec 24. Bioorg Med Chem. 2015. PMID: 25577708 Review.
Cited by
-
"Upcycling" known molecules and targets for drug-resistant TB.Front Cell Infect Microbiol. 2022 Oct 6;12:1029044. doi: 10.3389/fcimb.2022.1029044. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36275029 Free PMC article.
-
Elucidating the Antimycobacterial Mechanism of Action of Ciprofloxacin Using Metabolomics.Microorganisms. 2021 May 28;9(6):1158. doi: 10.3390/microorganisms9061158. Microorganisms. 2021. PMID: 34071153 Free PMC article.
References
-
- World Health Organisation . Global Tuberculosis Report 2017. http://apps.who.int/iris/bitstream/handle/10665/259366/9789241565516‐eng..., (2017) doi: WHO/HTM/TB/2017.23.
-
- World Health Organization . MDR TB/RR TB Factsheet TB 2017. in World Health Organization 1–2 (2017).
-
- Cole ST, Brosch R, Parkhill J, et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–544. - PubMed
-
- Hartmann G, Honikel KO, Knüsel F, Nüesch J. The specific inhibition of the DNA‐directed RNA synthesis by rifamycin. Biochim Biophys Acta. 1967;145:843–844. - PubMed
-
- Umezawa H, Mizuno S, Yamazaki H, Nitta K. Inhibition of DNA‐dependent RNA synthesis by rifamycins. J Antibiot. 1968;21:234–236. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
