Phage-DMS: A Comprehensive Method for Fine Mapping of Antibody Epitopes
- PMID: 33089110
- PMCID: PMC7566095
- DOI: 10.1016/j.isci.2020.101622
Phage-DMS: A Comprehensive Method for Fine Mapping of Antibody Epitopes
Abstract
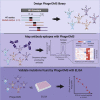
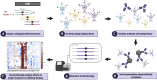
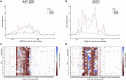
Understanding the antibody response is critical to developing vaccine and antibody-based therapies and has inspired the recent development of new methods to isolate antibodies. Methods to define the antibody-antigen interactions that determine specificity or allow escape have not kept pace. We developed Phage-DMS, a method that combines two powerful approaches-immunoprecipitation of phage peptide libraries and deep mutational scanning (DMS)-to enable high-throughput fine mapping of antibody epitopes. As an example, we designed sequences encoding all possible amino acid variants of HIV Envelope to create phage libraries. Using Phage-DMS, we identified sites of escape predicted using other approaches for four well-characterized HIV monoclonal antibodies with known linear epitopes. In some cases, the results of Phage-DMS refined the epitope beyond what was determined in previous studies. This method has the potential to rapidly and comprehensively screen many antibodies in a single experiment to define sites essential for binding interactions.
Keywords: Genomic Library; Virology.
© 2020 The Authors.
Conflict of interest statement
M.E.G. and J.O. are inventors on a patent application on Phage-DMS.
Figures



Similar articles
-
High-Resolution Mapping of Human Norovirus Antigens via Genomic Phage Display Library Selections and Deep Sequencing.J Virol. 2020 Dec 9;95(1):e01495-20. doi: 10.1128/JVI.01495-20. Print 2020 Dec 9. J Virol. 2020. PMID: 33055250 Free PMC article.
-
Comprehensive epitope analysis of monoclonal anti-proenkephalin antibodies using phage display libraries and synthetic peptides: revelation of antibody fine specificities caused by somatic mutations in the variable region genes.J Mol Biol. 1995 Apr 14;247(5):932-46. doi: 10.1006/jmbi.1995.0191. J Mol Biol. 1995. PMID: 7536850
-
Epitope mapping by phage display: random versus gene-fragment libraries.J Immunol Methods. 1997 Aug 7;206(1-2):43-52. doi: 10.1016/s0022-1759(97)00083-5. J Immunol Methods. 1997. PMID: 9328567
-
Epitope mapping via in vitro deep mutational scanning methods and its applications.J Biol Chem. 2025 Jan;301(1):108072. doi: 10.1016/j.jbc.2024.108072. Epub 2024 Dec 14. J Biol Chem. 2025. PMID: 39674321 Free PMC article. Review.
-
Epitope mapping: the first step in developing epitope-based vaccines.BioDrugs. 2007;21(3):145-56. doi: 10.2165/00063030-200721030-00002. BioDrugs. 2007. PMID: 17516710 Free PMC article. Review.
Cited by
-
Deep mutational scanning of proteins in mammalian cells.Cell Rep Methods. 2023 Nov 20;3(11):100641. doi: 10.1016/j.crmeth.2023.100641. Epub 2023 Nov 13. Cell Rep Methods. 2023. PMID: 37963462 Free PMC article. Review.
-
Elevated binding and functional antibody responses to SARS-CoV-2 in infants versus mothers.Nat Commun. 2023 Aug 11;14(1):4864. doi: 10.1038/s41467-023-40554-w. Nat Commun. 2023. PMID: 37567924 Free PMC article.
-
Comprehensive characterization of the antibody responses to SARS-CoV-2 Spike protein finds additional vaccine-induced epitopes beyond those for mild infection.Elife. 2022 Jan 24;11:e73490. doi: 10.7554/eLife.73490. Elife. 2022. PMID: 35072628 Free PMC article.
-
phippery: a software suite for PhIP-Seq data analysis.Bioinformatics. 2023 Oct 3;39(10):btad583. doi: 10.1093/bioinformatics/btad583. Bioinformatics. 2023. PMID: 37740324 Free PMC article.
-
Comprehensive characterization of the antibody responses to SARS-CoV-2 Spike protein after infection and/or vaccination.bioRxiv [Preprint]. 2021 Oct 5:2021.10.05.463210. doi: 10.1101/2021.10.05.463210. bioRxiv. 2021. Update in: Elife. 2022 Jan 24;11:e73490. doi: 10.7554/eLife.73490. PMID: 34642694 Free PMC article. Updated. Preprint.
References
-
- Cavacini L.A., Emes C.L., Wisnewski A.V., Power J., Lewis G., Montefiori D., Posner M.R. Functional and molecular characterization of human monoclonal antibody reactive with the immunodominant region of HIV type 1 glycoprotein 41. AIDS Res. Hum. Retroviruses. 1998;14:1271–1280. - PubMed
-
- Dall'Acqua W., Goldman E.R., Lin W., Teng C., Tsuchiya D., Li H., Ysern X., Braden B.C., Li Y., Smith-Gill S.J. A mutational analysis of binding interactions in an antigen-antibody protein-protein complex. Biochemistry. 1998;37:7981–7991. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

