Pharmacogenomics Biomarker Discovery and Validation for Translation in Clinical Practice
- PMID: 33089968
- PMCID: PMC7877857
- DOI: 10.1111/cts.12869
Pharmacogenomics Biomarker Discovery and Validation for Translation in Clinical Practice
Abstract
Interindividual variability in drug efficacy and toxicity is a major challenge in clinical practice. Variations in drug pharmacokinetics (PKs) and pharmacodynamics (PDs) can be, in part, explained by polymorphic variants in genes encoding drug metabolizing enzymes and transporters (absorption, distribution, metabolism, and excretion) or in genes encoding drug receptors. Pharmacogenomics (PGx) has allowed the identification of predictive biomarkers of drug PKs and PDs and the current knowledge of genome-disease and genome-drug interactions offers the opportunity to optimize tailored drug therapy. High-throughput PGx genotyping, from targeted to more comprehensive strategies, allows the identification of PK/PD genotypes to be developed as clinical predictive biomarkers. However, a biomarker needs a robust process of validation followed by clinical-grade assay development and must comply to stringent regulatory guidelines. We here discuss the methodological challenges and the emerging technological tools in PGx biomarker discovery and validation, at the crossroad among molecular genetics, bioinformatics, and clinical medicine.
© 2020 The Authors. Clinical and Translational Science published by Wiley Periodicals LLC on behalf of the American Society for Clinical Pharmacology and Therapeutics.
Conflict of interest statement
All authors declared no competing interests for this work.
Figures
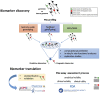
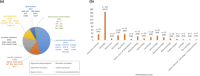
References
-
- Illing, P.T. , Purcell, A.W. & Mccluskey, J. The role of HLA genes in pharmacogenomics: unravelling HLA associated adverse drug reactions. Immunogenetics 69, 617–630 (2017). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

