Transcriptional profile of platelets and iPSC-derived megakaryocytes from whole-genome and RNA sequencing
- PMID: 33094331
- PMCID: PMC7918180
- DOI: 10.1182/blood.2020006115
Transcriptional profile of platelets and iPSC-derived megakaryocytes from whole-genome and RNA sequencing
Abstract
Genome-wide association studies have identified common variants associated with platelet-related phenotypes, but because these variants are largely intronic or intergenic, their link to platelet biology is unclear. In 290 normal subjects from the GeneSTAR Research Study (110 African Americans [AAs] and 180 European Americans [EAs]), we generated whole-genome sequence data from whole blood and RNA sequence data from extracted nonribosomal RNA from 185 induced pluripotent stem cell-derived megakaryocyte (MK) cell lines (platelet precursor cells) and 290 blood platelet samples from these subjects. Using eigenMT software to select the peak single-nucleotide polymorphism (SNP) for each expressed gene, and meta-analyzing the results of AAs and EAs, we identify (q-value < 0.05) 946 cis-expression quantitative trait loci (eQTLs) in derived MKs and 1830 cis-eQTLs in blood platelets. Among the 57 eQTLs shared between the 2 tissues, the estimated directions of effect are very consistent (98.2% concordance). A high proportion of detected cis-eQTLs (74.9% in MKs and 84.3% in platelets) are unique to MKs and platelets compared with peak-associated SNP-expressed gene pairs of 48 other tissue types that are reported in version V7 of the Genotype-Tissue Expression Project. The locations of our identified eQTLs are significantly enriched for overlap with several annotation tracks highlighting genomic regions with specific functionality in MKs, including MK-specific DNAse hotspots, H3K27-acetylation marks, H3K4-methylation marks, enhancers, and superenhancers. These results offer insights into the regulatory signature of MKs and platelets, with significant overlap in genes expressed, eQTLs detected, and enrichment within known superenhancers relevant to platelet biology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures
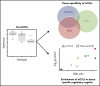


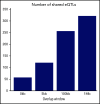

Comment in
-
eQTLs in platelets and iPSC-megakaryocytes.Blood. 2021 Feb 18;137(7):869-870. doi: 10.1182/blood.2020009461. Blood. 2021. PMID: 33599763 Free PMC article.
References
-
- Davì G, Patrono C. Platelet activation and atherothrombosis. N Engl J Med. 2007;357(24):2482-2494. - PubMed
-
- Colman R, Clowes A, George J, Hirsh J, Marder V. Overview of hemostasis. In: Colman RWHJ, Marder VJ, Clowes AW, George JN, eds. Hemostasis and Thrombosis Basic Principles and Practice, Philadelphia, PA: Lippincott Williams & Wilkins; 2001:3-16.
-
- Ashby B, Colman R, Daniel J, Kunapuli S, Smith J. Platelet stimulatory and inhibitory receptors. In: Colman RW, Hirsh J, Marder VJ, Clowes AW, George J, eds. Hemostasis and Thrombosis Basic Principles and Clinical Practice, Philadelphia, PA: Lippincott Williams & Wilkins; 2001:505-520.
-
- Abrams C, Brass L. Platelet signal transduction. In: Colman RW, Hirsh J, Marder VJ, Clowes AW, George J, eds. Hemostasis and Thrombosis Basic Principles and Clinical Practice. Philadelphia, PA: Lippincott Williams & Wilkins; 2001:541-559.
-
- Marcus AJ, Safier LB. Thromboregulation: multicellular modulation of platelet reactivity in hemostasis and thrombosis. FASEB J. 1993;7(6):516-522. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

