Premature aging syndrome showing random chromosome number instabilities with CDC20 mutation
- PMID: 33094908
- PMCID: PMC7681047
- DOI: 10.1111/acel.13251
Premature aging syndrome showing random chromosome number instabilities with CDC20 mutation
Abstract
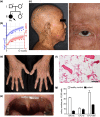
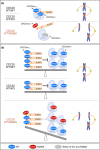
Damage to the genome can accelerate aging. The percentage of aneuploid cells, that is, cells with an abnormal number of chromosomes, increases during aging; however, it is not clear whether increased aneuploidy accelerates aging. Here, we report an individual showing premature aging phenotypes of various organs including early hair loss, atrophic skin, and loss of hematopoietic stem cells; instability of chromosome numbers known as mosaic variegated aneuploidy (MVA); and spindle assembly checkpoint (SAC) failure. Exome sequencing identified a de novo heterozygous germline missense mutation of c.856C>A (p.R286S) in the mitotic activator CDC20. The mutant CDC20 showed lower binding affinity to BUBR1 during the formation of the mitotic checkpoint complex (MCC), but not during the interaction between MCC and the anaphase-promoting complex/cyclosome (APC/C)-CDC20 complex. While heterozygous knockout of CDC20 did not induce SAC failure, knock-in of the mutant CDC20 induced SAC failure and random aneuploidy in cultured cells, indicating that the particular missense mutation is pathogenic probably via the resultant imbalance between MCC and APC/C-CDC20 complex. We postulate that accelerated chromosome number instability induces premature aging in humans, which may be associated with early loss of stem cells. These findings could form the basis of a novel disease model of the aging of the body and organs.
Keywords: Cdc20 proteins; M phase cell cycle checkpoints; aging; chromosomal instability; chromosome segregation; genomic instability; premature.
© 2020 The Authors. Aging Cell published by the Anatomical Society and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


Similar articles
-
Role of ubiquitylation of components of mitotic checkpoint complex in their dissociation from anaphase-promoting complex/cyclosome.Proc Natl Acad Sci U S A. 2018 Feb 20;115(8):1777-1782. doi: 10.1073/pnas.1720312115. Epub 2018 Feb 5. Proc Natl Acad Sci U S A. 2018. PMID: 29432156 Free PMC article.
-
High CDC20 levels increase sensitivity of cancer cells to MPS1 inhibitors.EMBO Rep. 2025 Feb;26(4):1036-1061. doi: 10.1038/s44319-024-00363-8. Epub 2025 Jan 21. EMBO Rep. 2025. PMID: 39838194 Free PMC article.
-
The Bub1-Plk1 kinase complex promotes spindle checkpoint signalling through Cdc20 phosphorylation.Nat Commun. 2016 Feb 25;7:10818. doi: 10.1038/ncomms10818. Nat Commun. 2016. PMID: 26912231 Free PMC article.
-
Cdc20: At the Crossroads between Chromosome Segregation and Mitotic Exit.Trends Biochem Sci. 2017 Mar;42(3):193-205. doi: 10.1016/j.tibs.2016.12.001. Epub 2017 Feb 12. Trends Biochem Sci. 2017. PMID: 28202332 Review.
-
BubR1 kinase: protection against aneuploidy and premature aging.Trends Mol Med. 2015 Jun;21(6):364-72. doi: 10.1016/j.molmed.2015.04.003. Epub 2015 May 8. Trends Mol Med. 2015. PMID: 25964054 Review.
Cited by
-
CDC20 in and out of mitosis: a prognostic factor and therapeutic target in hematological malignancies.J Exp Clin Cancer Res. 2022 Apr 30;41(1):159. doi: 10.1186/s13046-022-02363-9. J Exp Clin Cancer Res. 2022. PMID: 35490245 Free PMC article. Review.
-
NBS1 I171V variant underlies individual differences in chromosomal radiosensitivity within human populations.Sci Rep. 2021 Oct 4;11(1):19661. doi: 10.1038/s41598-021-98673-7. Sci Rep. 2021. PMID: 34608183 Free PMC article.
-
Mosaic variegated aneuploidy in development, ageing and cancer.Nat Rev Genet. 2024 Dec;25(12):864-878. doi: 10.1038/s41576-024-00762-6. Epub 2024 Aug 21. Nat Rev Genet. 2024. PMID: 39169218 Review.
-
The Anaphase-Promoting Complex/Cyclosome Is a Cellular Ageing Regulator.Int J Mol Sci. 2022 Dec 5;23(23):15327. doi: 10.3390/ijms232315327. Int J Mol Sci. 2022. PMID: 36499653 Free PMC article. Review.
-
CDC20-Mediated Selective Autophagy Degradation of PBRM1 Affects Immunotherapy for Renal Cell Carcinoma.Adv Sci (Weinh). 2025 Feb;12(5):e2412967. doi: 10.1002/advs.202412967. Epub 2024 Dec 10. Adv Sci (Weinh). 2025. PMID: 39656940 Free PMC article.
References
-
- Baker, D. J. , Jeganathan, K. B. , Cameron, J. D. , Thompson, M. , Juneja, S. , Kopecka, A. , Kumar, R. , Jenkins, R. B. , de Groen, P. C. , Roche, P. , & van Deursen, J. M. (2004). BubR1 insufficiency causes early onset of aging‐associated phenotypes and infertility in mice. Nature Genetics, 36(7), 744–749. 10.1038/ng1382 - DOI - PubMed
-
- Barbaux, S. , Niaudet, P. , Gubler, M.‐C. , Grünfeld, J.‐P. , Jaubert, F. , Kuttenn, F. , Fékété, C. N. , Souleyreau‐Therville, N. , Thibaud, E. , Fellous, M. , & McElreavey, K. (1997). Donor splice‐site mutations in WT1 are responsible for Frasier syndrome. Nature Genetics, 17(4), 467–470. 10.1038/ng1297-467 - DOI - PubMed
-
- Database of Single Nucleotide Polymorphisms (dbSNP) (2019). Bethesda (MD): National Center for Biotechnology Information, National Library of Medicine. dbSNP gene ID:991, (dbSNP Build ID: 153 (Aug 8, 2019)). Retrieved from https://www.ncbi.nlm.nih.gov/SNP/snp_ref.cgi?locusId=991
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

