Identification of Specific Circular RNA Expression Patterns and MicroRNA Interaction Networks in Mesial Temporal Lobe Epilepsy
- PMID: 33101384
- PMCID: PMC7546880
- DOI: 10.3389/fgene.2020.564301
Identification of Specific Circular RNA Expression Patterns and MicroRNA Interaction Networks in Mesial Temporal Lobe Epilepsy
Abstract
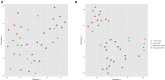
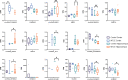
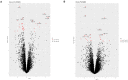
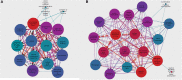
Circular RNAs (circRNAs) regulate mRNA translation by binding to microRNAs (miRNAs), and their expression is altered in diverse disorders, including cancer, cardiovascular disease, and Parkinson's disease. Here, we compare circRNA expression patterns in the temporal cortex and hippocampus of patients with pharmacoresistant mesial temporal lobe epilepsy (MTLE) and healthy controls. Nine circRNAs showed significant differential expression, including circRNA-HOMER1, which is expressed in synapses. Further, we identified miRNA binding sites within the sequences of differentially expressed (DE) circRNAs; expression levels of mRNAs correlated with changes in complementary miRNAs. Gene set enrichment analysis of mRNA targets revealed functions in heterocyclic compound binding, regulation of transcription, and signal transduction, which maintain the structure and function of hippocampal neurons. The circRNA-miRNA-mRNA interaction networks illuminate the molecular changes in MTLE, which may be pathogenic or an effect of the disease or treatments and suggests that DE circRNAs and associated miRNAs may be novel therapeutic targets.
Keywords: RNA-Seq; circular RNAs; epilepsy; gene expression; mesial temporal lobe epilepsy; microRNAs.
Copyright © 2020 Gray, Mills, Curry-Hyde, Devore, Friedman, Thom, Scott, Thijs, Aronica, Devinsky and Janitz.
Figures
Similar articles
-
Expression of Circ_Satb1 Is Decreased in Mesial Temporal Lobe Epilepsy and Regulates Dendritic Spine Morphology.Front Mol Neurosci. 2022 Mar 3;15:832133. doi: 10.3389/fnmol.2022.832133. eCollection 2022. Front Mol Neurosci. 2022. PMID: 35310884 Free PMC article.
-
MicroRNA and mesial temporal lobe epilepsy with hippocampal sclerosis: Whole miRNome profiling of human hippocampus.Epilepsia. 2017 Oct;58(10):1782-1793. doi: 10.1111/epi.13870. Epub 2017 Aug 16. Epilepsia. 2017. PMID: 28815576
-
Possible epigenetic regulatory effect of dysregulated circular RNAs in epilepsy.PLoS One. 2018 Dec 28;13(12):e0209829. doi: 10.1371/journal.pone.0209829. eCollection 2018. PLoS One. 2018. PMID: 30592747 Free PMC article.
-
Crosstalk in competing endogenous RNA networks reveals new circular RNAs involved in the pathogenesis of early HIV infection.J Transl Med. 2018 Nov 29;16(1):332. doi: 10.1186/s12967-018-1706-1. J Transl Med. 2018. PMID: 30486834 Free PMC article.
-
MicroRNAs in epilepsy: pathophysiology and clinical utility.Lancet Neurol. 2016 Dec;15(13):1368-1376. doi: 10.1016/S1474-4422(16)30246-0. Lancet Neurol. 2016. PMID: 27839653 Review.
Cited by
-
Identification and characterization of circular RNA in the model of autism spectrum disorder from PM2.5 exposure.Front Genet. 2023 May 9;14:970465. doi: 10.3389/fgene.2023.970465. eCollection 2023. Front Genet. 2023. PMID: 37229188 Free PMC article.
-
The Coordination of mTOR Signaling and Non-Coding RNA in Regulating Epileptic Neuroinflammation.Front Immunol. 2022 Jul 11;13:924642. doi: 10.3389/fimmu.2022.924642. eCollection 2022. Front Immunol. 2022. PMID: 35898503 Free PMC article. Review.
-
CircRNA-associated ceRNA regulatory networks as emerging mechanisms governing the development and biophysiopathology of epilepsy.CNS Neurosci Ther. 2024 Apr;30(4):e14735. doi: 10.1111/cns.14735. CNS Neurosci Ther. 2024. PMID: 38676299 Free PMC article. Review.
-
Multi-omic strategies applied to the study of pharmacoresistance in mesial temporal lobe epilepsy.Epilepsia Open. 2022 Aug;7 Suppl 1(Suppl 1):S94-S120. doi: 10.1002/epi4.12536. Epub 2021 Oct 18. Epilepsia Open. 2022. PMID: 34486831 Free PMC article. Review.
-
Expression of Circ_Satb1 Is Decreased in Mesial Temporal Lobe Epilepsy and Regulates Dendritic Spine Morphology.Front Mol Neurosci. 2022 Mar 3;15:832133. doi: 10.3389/fnmol.2022.832133. eCollection 2022. Front Mol Neurosci. 2022. PMID: 35310884 Free PMC article.
References
-
- Andrews S. (2018). FastQC a quality control tool for high throughput sequence data. Babraham. Bioinfo. 2018 3–5.
-
- Blümcke I., Thom M., Aronica E., Armstrong D. D., Bartolomei F., Bernasconi A., et al. (2013). International consensus classification of hippocampal sclerosis in temporal lobe epilepsy: A Task Force report from the ILAE Commission on Diagnostic Methods. Epilepsia 54 1315–1329. 10.1111/epi.12220 - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

