Hybrid MM/CG Webserver: Automatic Set Up of Molecular Mechanics/Coarse-Grained Simulations for Human G Protein-Coupled Receptor/Ligand Complexes
- PMID: 33102525
- PMCID: PMC7500467
- DOI: 10.3389/fmolb.2020.576689
Hybrid MM/CG Webserver: Automatic Set Up of Molecular Mechanics/Coarse-Grained Simulations for Human G Protein-Coupled Receptor/Ligand Complexes
Abstract
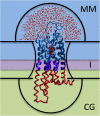
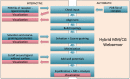
Hybrid Molecular Mechanics/Coarse-Grained (MM/CG) simulations help predict ligand poses in human G protein-coupled receptors (hGPCRs), the most important protein superfamily for pharmacological applications. This approach allows the description of the ligand, the binding cavity, and the surrounding water molecules at atomistic resolution, while coarse-graining the rest of the receptor. Here, we present the Hybrid MM/CG Webserver (mmcg.grs.kfa-juelich.de) that automatizes and speeds up the MM/CG simulation setup of hGPCR/ligand complexes. Initial structures for such complexes can be easily and efficiently generated with other webservers. The Hybrid MM/CG server also allows for equilibration of the systems, either fully automatically or interactively. The results are visualized online (using both interactive 3D visualizations and analysis plots), helping the user identify possible issues and modify the setup parameters accordingly. Furthermore, the prepared system can be downloaded and the simulation continued locally.
Keywords: G protein-coupled receptor; MM/CG; coarse-grained; hybrid methods; ligand; molecular dynamics simulation; molecular mechanics; webserver.
Copyright © 2020 Schneider, Ribeiro, Alfonso-Prieto, Carloni and Giorgetti.
Figures

Similar articles
-
Ligand Pose Predictions for Human G Protein-Coupled Receptors: Insights from the Amber-Based Hybrid Molecular Mechanics/Coarse-Grained Approach.J Chem Inf Model. 2020 Oct 26;60(10):5103-5116. doi: 10.1021/acs.jcim.0c00661. Epub 2020 Aug 14. J Chem Inf Model. 2020. PMID: 32786708
-
Hybrid simulations: combining atomistic and coarse-grained force fields using virtual sites.Phys Chem Chem Phys. 2011 Jun 14;13(22):10437-48. doi: 10.1039/c0cp02981e. Epub 2011 Apr 15. Phys Chem Chem Phys. 2011. PMID: 21494747
-
Hybrid molecular mechanics/coarse-grained simulations for structural prediction of G-protein coupled receptor/ligand complexes.PLoS One. 2012;7(10):e47332. doi: 10.1371/journal.pone.0047332. Epub 2012 Oct 19. PLoS One. 2012. PMID: 23094046 Free PMC article.
-
Predicting ligand binding poses for low-resolution membrane protein models: Perspectives from multiscale simulations.Biochem Biophys Res Commun. 2018 Mar 29;498(2):366-374. doi: 10.1016/j.bbrc.2018.01.160. Epub 2018 Feb 2. Biochem Biophys Res Commun. 2018. PMID: 29409902 Review.
-
Understanding Ligand Binding to G-Protein Coupled Receptors Using Multiscale Simulations.Front Mol Biosci. 2019 May 3;6:29. doi: 10.3389/fmolb.2019.00029. eCollection 2019. Front Mol Biosci. 2019. PMID: 31131282 Free PMC article. Review.
Cited by
-
GPCRsignal: webserver for analysis of the interface between G-protein-coupled receptors and their effector proteins by dynamics and mutations.Nucleic Acids Res. 2021 Jul 2;49(W1):W247-W256. doi: 10.1093/nar/gkab434. Nucleic Acids Res. 2021. PMID: 34060630 Free PMC article.
-
From System Modeling to System Analysis: The Impact of Resolution Level and Resolution Distribution in the Computer-Aided Investigation of Biomolecules.Front Mol Biosci. 2021 Jun 7;8:676976. doi: 10.3389/fmolb.2021.676976. eCollection 2021. Front Mol Biosci. 2021. PMID: 34164432 Free PMC article. Review.
-
CGMD Platform: Integrated Web Servers for the Preparation, Running, and Analysis of Coarse-Grained Molecular Dynamics Simulations.Molecules. 2020 Dec 15;25(24):5934. doi: 10.3390/molecules25245934. Molecules. 2020. PMID: 33333836 Free PMC article.
References
-
- Berendsen H. J. C., van der Spoel D., van Drunen R. (1995). GROMACS: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91 43–56. 10.1016/0010-4655(95)00042-e - DOI
LinkOut - more resources
Full Text Sources

