A curated list of genes that affect the plant ionome
- PMID: 33103043
- PMCID: PMC7576880
- DOI: 10.1002/pld3.272
A curated list of genes that affect the plant ionome
Abstract
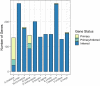
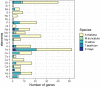
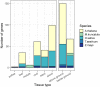
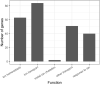
Understanding the mechanisms underlying plants' adaptation to their environment will require knowledge of the genes and alleles underlying elemental composition. Modern genetics is capable of quickly, and cheaply indicating which regions of DNA are associated with particular phenotypes in question, but most genes remain poorly annotated, hindering the identification of candidate genes. To help identify candidate genes underlying elemental accumulations, we have created the known ionome gene (KIG) list: a curated collection of genes experimentally shown to change uptake, accumulation, and distribution of elements. We have also created an automated computational pipeline to generate lists of KIG orthologs in other plant species using the PhytoMine database. The current version of KIG consists of 176 known genes covering 5 species, 23 elements, and their 1588 orthologs in 10 species. Analysis of the known genes demonstrated that most were identified in the model plant Arabidopsis thaliana, and that transporter coding genes and genes altering the accumulation of iron and zinc are overrepresented in the current list.
Keywords: curated; ionomics; mineral nutrition.
© 2020 The Authors. Plant Direct published by American Society of Plant Biologists, Society for Experimental Biology and John Wiley & Sons Ltd.
Figures
Similar articles
-
Plant Ionomics: From Elemental Profiling to Environmental Adaptation.Mol Plant. 2016 Jun 6;9(6):787-97. doi: 10.1016/j.molp.2016.05.003. Epub 2016 May 19. Mol Plant. 2016. PMID: 27212388 Review.
-
Should we treat the ionome as a combination of individual elements, or should we be deriving novel combined traits?J Exp Bot. 2015 Apr;66(8):2127-31. doi: 10.1093/jxb/erv040. Epub 2015 Feb 24. J Exp Bot. 2015. PMID: 25711709 Free PMC article. Review.
-
Ionomics and the study of the plant ionome.Annu Rev Plant Biol. 2008;59:709-33. doi: 10.1146/annurev.arplant.59.032607.092942. Annu Rev Plant Biol. 2008. PMID: 18251712 Review.
-
1,135 ionomes reveal the global pattern of leaf and seed mineral nutrient and trace element diversity in Arabidopsis thaliana.Plant J. 2021 Apr;106(2):536-554. doi: 10.1111/tpj.15177. Epub 2021 Mar 23. Plant J. 2021. PMID: 33506585
-
Ionomics: studying the social network of mineral nutrients.Curr Opin Plant Biol. 2009 Jun;12(3):381-6. doi: 10.1016/j.pbi.2009.05.002. Epub 2009 May 28. Curr Opin Plant Biol. 2009. PMID: 19481970 Free PMC article. Review.
Cited by
-
Drought specifically downregulates mineral nutrition: Plant ionomic content and associated gene expression.Plant Direct. 2022 Aug 5;6(8):e402. doi: 10.1002/pld3.402. eCollection 2022 Aug. Plant Direct. 2022. PMID: 35949952 Free PMC article.
-
Ionomic Approaches for Discovery of Novel Stress-Resilient Genes in Plants.Int J Mol Sci. 2021 Jul 2;22(13):7182. doi: 10.3390/ijms22137182. Int J Mol Sci. 2021. PMID: 34281232 Free PMC article. Review.
-
The vacuolar H+/Ca transporter CAX1 participates in submergence and anoxia stress responses.Plant Physiol. 2022 Nov 28;190(4):2617-2636. doi: 10.1093/plphys/kiac375. Plant Physiol. 2022. PMID: 35972350 Free PMC article.
-
Specificity and Plasticity of the Functional Ionome of Brassica napus and Triticum aestivum Exposed to Micronutrient or Beneficial Nutrient Deprivation and Predictive Sensitivity of the Ionomic Signatures.Front Plant Sci. 2021 Feb 10;12:641678. doi: 10.3389/fpls.2021.641678. eCollection 2021. Front Plant Sci. 2021. PMID: 33643368 Free PMC article.
-
Iron Biofortification in Rice: An Update on Quantitative Trait Loci and Candidate Genes.Front Plant Sci. 2021 May 26;12:647341. doi: 10.3389/fpls.2021.647341. eCollection 2021. Front Plant Sci. 2021. PMID: 34122472 Free PMC article. Review.
References
-
- Abreu, I. , Saéz, Á. , Castro‐Rodríguez, R. , Escudero, V. , Rodríguez‐Haas, B. , Senovilla, M. , Larue, C. , Grolimund, D. , Tejada‐Jiménez, M. , Imperial, J. , & González‐Guerrero, M. (2017). Medicago truncatula zinc‐iron permease6 provides zinc to rhizobia‐infected nodule cells. Plant, Cell & Environment, 40(11), 2706–2719. - PubMed
-
- Aggarwal, S. , Kumar, A. , Bhati, K. K. , Kaur, G. , Shukla, V. , Tiwari, S. , & Pandey, A. K. (2018). RNAi‐mediated downregulation of inositol pentakisphosphate kinase (IPK1) in wheat grains decreases phytic acid levels and increases Fe and Zn Accumulation. Frontiers in Plant Science, 9(March), 259 10.3389/fpls.2018.00259 - DOI - PMC - PubMed
-
- Ahmad, I. , Devonshire, J. , Mohamed, R. , Schultze, M. , & Maathuis, F. J. M. (2016). Overexpression of the potassium channel TPKb in small vacuoles confers osmotic and drought tolerance to rice. The New Phytologist, 209(3), 1040–1048. - PubMed
-
- Andrés‐Colás, N. , Sancenón, V. , Rodríguez‐Navarro, S. , Mayo, S. , Thiele, D. J. , Ecker, J. R. , Puig, S. , & Peñarrubia, L. (2006). The arabidopsis heavy metal P‐type ATPase HMA5 interacts with metallochaperones and functions in copper detoxification of roots. The Plant Journal, 45(2), 225–236. 10.1111/j.1365-313X.2005.02601.x - DOI - PubMed
LinkOut - more resources
Full Text Sources

