Comprehensive Analysis of DNA Methylation and Prediction of Response to NeoadjuvantTherapy in Locally Advanced Rectal Cancer
- PMID: 33105711
- PMCID: PMC7690383
- DOI: 10.3390/cancers12113079
Comprehensive Analysis of DNA Methylation and Prediction of Response to NeoadjuvantTherapy in Locally Advanced Rectal Cancer
Abstract
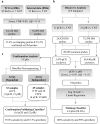
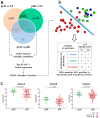
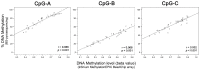
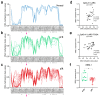
The treatment for locally advanced rectal carcinomas (LARC) is based on neoadjuvant chemoradiotherapy (nCRT) and surgery, which results in pathological complete response (pCR) in up to 30% of patients. Since epigenetic changes may influence response to therapy, we aimed to identify DNA methylation markers predictive of pCR in LARC patients treated with nCRT. We used high-throughput DNA methylation analysis of 32 treatment-naïve LARC biopsies and five normal rectal tissues to explore the predictive value of differentially methylated (DM) CpGs. External validation was carried out with The Cancer Genome Atlas-Rectal Adenocarcinoma (TCGA-READ 99 cases). A classifier based on three-CpGs DM (linked to OBSL1, GPR1, and INSIG1 genes) was able to discriminate pCR from incomplete responders with high sensitivity and specificity. The methylation levels of the selected CpGs confirmed the predictive value of our classifier in 77 LARCs evaluated by bisulfite pyrosequencing. Evaluation of external datasets (TCGA-READ, GSE81006, GSE75546, and GSE39958) reproduced our results. As the three CpGs were mapped near to regulatory elements, we performed an integrative analysis in regions associated with predicted cis-regulatory elements. A positive and inverse correlation between DNA methylation and gene expression was found in two CpGs. We propose a novel predictive tool based on three CpGs potentially useful for pretreatment screening of LARC patients and guide the selection of treatment modality.
Keywords: 5-fluorouracil; high-throughput DNA methylation analysis; predictive biomarker; translational research.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Tevis S.E., Kohlnhofer B.M., Stringfield S., Foley E.F., Harms B.A., Heise C.P., Kennedy G.D. Postoperative complications in patients with rectal cancer are associated with delays in chemotherapy that lead to worse disease-free and overall survival. Dis. Colon Rectum. 2013;56:1339–1348. doi: 10.1097/DCR.0b013e3182a857eb. - DOI - PMC - PubMed
-
- Giandomenico F., Gavaruzzi T., Lotto L., Del Bianco P., Barina A., Perin A., Pucciarelli S. Quality of life after surgery for rectal cancer: A systematic review of comparisons with the general population. Expert Rev. Gastroenterol. Hepatol. 2015;9:1227–1242. doi: 10.1586/17474124.2015.1070667. - DOI - PubMed
-
- Fokas E., Liersch T., Fietkau R., Hohenberger W., Beissbarth T., Hess C., Becker H., Ghadimi M., Mrak K., Merkel S., et al. Tumor regression grading after preoperative chemoradiotherapy for locally advanced rectal carcinoma revisited: Updated results of the CAO/ARO/AIO-94 trial. J. Clin. Oncol. 2014;32:1554–1562. doi: 10.1200/JCO.2013.54.3769. - DOI - PubMed
-
- Lorimer P.D., Motz B.M., Kirks R.C., Boselli D.M., Walsh K.K., Prabhu R.S., Hill J.S., Salo J.C. Pathologic Complete Response Rates After Neoadjuvant Treatment in Rectal Cancer: An Analysis of the National Cancer Database. Ann. Surg. Oncol. 2017;24:2095–2103. doi: 10.1245/s10434-017-5873-8. - DOI - PubMed
Grants and funding
- 2008/57887-9/Fundação de Amparo à Pesquisa do Estado de São Paulo
- 2014/06323-9/Fundação de Amparo à Pesquisa do Estado de São Paulo
- 2015/25803-4/Fundação de Amparo à Pesquisa do Estado de São Paulo
- 573589/08-9/Conselho Nacional de Desenvolvimento Científico e Tecnológico
- #/Danish Colorectal Cancer Center South
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

