Inhibition of Drp1 SUMOylation by ALR protects the liver from ischemia-reperfusion injury
- PMID: 33110216
- PMCID: PMC8027887
- DOI: 10.1038/s41418-020-00641-7
Inhibition of Drp1 SUMOylation by ALR protects the liver from ischemia-reperfusion injury
Abstract
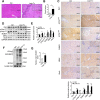
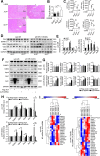
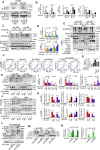
Hepatic ischemic reperfusion injury (IRI) is a common complication of liver surgery. Although an imbalance between mitochondrial fission and fusion has been identified as the cause of IRI, the detailed mechanism remains unclear. Augmenter of liver regeneration (ALR) was reported to prevent mitochondrial fission by inhibiting dynamin-related protein 1 (Drp1) phosphorylation, contributing partially to its liver protection. Apart from phosphorylation, Drp1 activity is also regulated by small ubiquitin-like modification (SUMOylation), which accelerates mitochondrial fission. This study aimed to investigate whether ALR-mediated protection from hepatic IRI might be associated with an effect on Drp1 SUMOylation. Liver tissues were harvested from both humans and from heterozygous ALR knockout mice, which underwent IRI. The SUMOylation and phosphorylation of Drp1 and their modulation by ALR were investigated. Hepatic Drp1 SUMOylation was significantly increased in human transplanted livers and IRI-livers of mice. ALR-transfection significantly decreased Drp1 SUMOylation, attenuated the IRI-induced mitochondrial fission and preserved mitochondrial stability and function. This study showed that the binding of transcription factor Yin Yang-1 (YY1) to its downstream target gene UBA2, a subunit of SUMO-E1 enzyme heterodimer, was critical to control Drp1 SUMOylation. By interacting with YY1, ALR inhibits its nuclear import and dramatically decreases the transcriptional level of UBA2. Consequently, mitochondrial fission was significantly reduced, and mitochondrial function was maintained. This study showed that the regulation of Drp1 SUMOylation by ALR protects mitochondria from fission, rescuing hepatocytes from IRI-induced apoptosis. These new findings provide a potential target for clinical intervention to reduce the effects of IRI during hepatic surgery.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





References
-
- Huguet C, Addario-Chieco P, Gavelli A, Arrigo E, Harb J, Clement RR. Technique of hepatic vascular exclusion for extensive liver resection. Am J Surg. 1992;163:602–5. - PubMed
-
- Abdalla EK, Roger N, Jacques B. Hepatic vascular occlusion: which technique? Surg Clin North Am. 2004;84:563–85. - PubMed
-
- Selzner M, Rüdiger HA, Sindram D, Madden J, Clavien PA. Mechanisms of ischemic injury are different in the steatotic and normal rat liver. Hepatology. 2000;32:1280–8. - PubMed
-
- Odeh M. The role of reperfusion-induced injury in the pathogenesis of the crush syndrome. N. Engl J Med. 1991;324:1417–22. - PubMed
-
- Elias-Miró M, Jiménez-Castro MB, Rodés J, Peralta C. Current knowledge on oxidative stress in hepatic ischemia/reperfusion. Free Radic Res. 2013;47:555–68. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

