Calcium dynamics and chromatin remodelling underlie heterogeneity in prolactin transcription
- PMID: 33112804
- PMCID: PMC7774774
- DOI: 10.1530/JME-20-0223
Calcium dynamics and chromatin remodelling underlie heterogeneity in prolactin transcription
Abstract
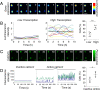
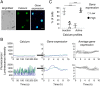
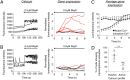
Pituitary cells have been reported to show spontaneous calcium oscillations and dynamic transcription cycles. To study both processes in the same living cell in real time, we used rat pituitary GH3 cells stably expressing human prolactin-luciferase or prolactin-EGFP reporter gene constructs loaded with a fluorescent calcium indicator and measured activity using single-cell time-lapse microscopy. We observed heterogeneity between clonal cells in the calcium activity and prolactin transcription in unstimulated conditions. There was a significant correlation between cells displaying spontaneous calcium spikes and cells showing spontaneous bursts in prolactin expression. Notably, cells showing no basal calcium activity showed low prolactin expression but elicited a significantly greater transcriptional response to BayK8644 compared to cells showing basal calcium activity. This suggested the presence of two subsets of cells within the population at any one time. Fluorescence-activated cell sorting was used to sort cells into two populations based on the expression level of prolactin-EGFP however, the bimodal pattern of expression was restored within 26 h. Chromatin immunoprecipitation showed that these sorted populations were distinct due to the extent of histone acetylation. We suggest that maintenance of a heterogeneous bimodal population is a fundamental characteristic of this cell type and that calcium activation and histone acetylation, at least in part, drive prolactin transcriptional competence.
Keywords: calcium; chromatin; heterogeneity; prolactin; single cell; transcription.
Figures



Similar articles
-
Dynamic analysis of stochastic transcription cycles.PLoS Biol. 2011 Apr;9(4):e1000607. doi: 10.1371/journal.pbio.1000607. Epub 2011 Apr 12. PLoS Biol. 2011. PMID: 21532732 Free PMC article.
-
Insulin-activated Elk-1 recruits the TIP60/NuA4 complex to increase prolactin gene transcription.Mol Cell Endocrinol. 2014 Jan 25;382(1):159-169. doi: 10.1016/j.mce.2013.09.021. Epub 2013 Sep 26. Mol Cell Endocrinol. 2014. PMID: 24075908
-
Epigenetic mechanisms in the dopamine D2 receptor-dependent inhibition of the prolactin gene.Mol Endocrinol. 2005 Jul;19(7):1904-17. doi: 10.1210/me.2004-0111. Epub 2005 Feb 24. Mol Endocrinol. 2005. PMID: 15731170
-
The prolactin gene: a paradigm of tissue-specific gene regulation with complex temporal transcription dynamics.J Neuroendocrinol. 2012 Jul;24(7):977-90. doi: 10.1111/j.1365-2826.2012.02310.x. J Neuroendocrinol. 2012. PMID: 22420298 Free PMC article. Review.
-
Histone acetylation and methylation: combinatorial players for transcriptional regulation.Subcell Biochem. 2007;41:351-69. Subcell Biochem. 2007. PMID: 17484136 Review.
Cited by
-
Establishment and Maintenance of Heat-Stress Memory in Plants.Int J Mol Sci. 2024 Aug 18;25(16):8976. doi: 10.3390/ijms25168976. Int J Mol Sci. 2024. PMID: 39201662 Free PMC article. Review.
-
Hypoxia-induced epigenetic regulation of breast cancer progression and the tumour microenvironment.Front Cell Dev Biol. 2024 Aug 30;12:1421629. doi: 10.3389/fcell.2024.1421629. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39282472 Free PMC article. Review.
References
-
- Charles AC, Piros ET, Evans CJ, Hales TG.1999. L-type Ca2+ channels and K+ channels specifically modulate the frequency and amplitude of spontaneous Ca2+ oscillations and have distinct roles in prolactin release in GH3 cells. Journal of Biological Chemistry 274 7508–7515. (10.1074/jbc.274.11.7508) - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

