Coronavirus biology and replication: implications for SARS-CoV-2
- PMID: 33116300
- PMCID: PMC7592455
- DOI: 10.1038/s41579-020-00468-6
Coronavirus biology and replication: implications for SARS-CoV-2
Abstract
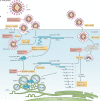
The SARS-CoV-2 pandemic and its unprecedented global societal and economic disruptive impact has marked the third zoonotic introduction of a highly pathogenic coronavirus into the human population. Although the previous coronavirus SARS-CoV and MERS-CoV epidemics raised awareness of the need for clinically available therapeutic or preventive interventions, to date, no treatments with proven efficacy are available. The development of effective intervention strategies relies on the knowledge of molecular and cellular mechanisms of coronavirus infections, which highlights the significance of studying virus-host interactions at the molecular level to identify targets for antiviral intervention and to elucidate critical viral and host determinants that are decisive for the development of severe disease. In this Review, we summarize the first discoveries that shape our current understanding of SARS-CoV-2 infection throughout the intracellular viral life cycle and relate that to our knowledge of coronavirus biology. The elucidation of similarities and differences between SARS-CoV-2 and other coronaviruses will support future preparedness and strategies to combat coronavirus infections.
Conflict of interest statement
The authors declare no competing interests.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

