Genetic-Based Hypertension Subtype Identification Using Informative SNPs
- PMID: 33121163
- PMCID: PMC7693873
- DOI: 10.3390/genes11111265
Genetic-Based Hypertension Subtype Identification Using Informative SNPs
Abstract
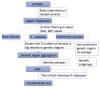
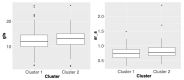
In this work, we proposed a process to select informative genetic variants for identifying clinically meaningful subtypes of hypertensive patients. We studied 575 African American (AA) and 612 Caucasian hypertensive participants enrolled in the Hypertension Genetic Epidemiology Network (HyperGEN) study and analyzed each race-based group separately. All study participants underwent GWAS (Genome-Wide Association Studies) and echocardiography. We applied a variety of statistical methods and filtering criteria, including generalized linear models, F statistics, burden tests, deleterious variant filtering, and others to select the most informative hypertension-related genetic variants. We performed an unsupervised learning algorithm non-negative matrix factorization (NMF) to identify hypertension subtypes with similar genetic characteristics. Kruskal-Wallis tests were used to demonstrate the clinical meaningfulness of genetic-based hypertension subtypes. Two subgroups were identified for both African American and Caucasian HyperGEN participants. In both AAs and Caucasians, indices of cardiac mechanics differed significantly by hypertension subtypes. African Americans tend to have more genetic variants compared to Caucasians; therefore, using genetic information to distinguish the disease subtypes for this group of people is relatively challenging, but we were able to identify two subtypes whose cardiac mechanics have statistically different distributions using the proposed process. The research gives a promising direction in using statistical methods to select genetic information and identify subgroups of diseases, which may inform the development and trial of novel targeted therapies.
Keywords: NMF; clustering algorithm; hypertension; subtype identification; variable selection.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Variants on chromosome 6p22.3 associated with blood pressure in the HyperGEN study: follow-up of FBPP quantitative trait loci.Am J Hypertens. 2011 Nov;24(11):1227-33. doi: 10.1038/ajh.2011.140. Epub 2011 Aug 18. Am J Hypertens. 2011. PMID: 21850057 Free PMC article.
-
Sequence variation of bradykinin receptors B1 and B2 and association with hypertension.J Hypertens. 2005 Jan;23(1):55-62. doi: 10.1097/00004872-200501000-00013. J Hypertens. 2005. PMID: 15643125
-
Haplotype association analysis of AGT variants with hypertension-related traits: the HyperGEN study.Hum Hered. 2005;60(3):164-76. doi: 10.1159/000090118. Epub 2005 Dec 12. Hum Hered. 2005. PMID: 16352906
-
Genetics of hypertension and cardiovascular disease and their interconnected pathways: lessons from large studies.Curr Hypertens Rep. 2011 Feb;13(1):46-54. doi: 10.1007/s11906-010-0174-7. Curr Hypertens Rep. 2011. PMID: 21128019 Free PMC article. Review.
-
A review of genetics, arterial stiffness, and blood pressure in African Americans.J Cardiovasc Transl Res. 2012 Jun;5(3):302-8. doi: 10.1007/s12265-012-9362-y. Epub 2012 Apr 11. J Cardiovasc Transl Res. 2012. PMID: 22492025 Free PMC article. Review.
Cited by
-
Hypertension in African Populations: Review and Computational Insights.Genes (Basel). 2021 Apr 6;12(4):532. doi: 10.3390/genes12040532. Genes (Basel). 2021. PMID: 33917487 Free PMC article. Review.
-
Northwestern University resource and education development initiatives to advance collaborative artificial intelligence across the learning health system.Learn Health Syst. 2024 Apr 15;8(3):e10417. doi: 10.1002/lrh2.10417. eCollection 2024 Jul. Learn Health Syst. 2024. PMID: 39036530 Free PMC article.
-
Unsupervised clustering analysis of SARS-Cov-2 population structure reveals six major subtypes at early stage across the world.bioRxiv [Preprint]. 2021 Nov 24:2020.09.04.283358. doi: 10.1101/2020.09.04.283358. bioRxiv. 2021. PMID: 34845455 Free PMC article. Preprint.
-
Multi-Trait Genetic Analysis Reveals Clinically Interpretable Hypertension Subtypes.Circ Genom Precis Med. 2022 Aug;15(4):e003583. doi: 10.1161/CIRCGEN.121.003583. Epub 2022 May 23. Circ Genom Precis Med. 2022. PMID: 35604428 Free PMC article.
References
-
- Farasat S.M., Morrell C.H., Scuteri A., Ting C.T., Yin C.P., Spurgeon H.A., Chen C.-H., Lakatta E.G., Najjar S.S. Do hypertensive individuals have enlarged aortic root diameters? Insights from studying the various subtypes of hypertension. Am. J. Hypertens. 2008;21:558–563. doi: 10.1038/ajh.2008.10. - DOI - PMC - PubMed
-
- Gupta R.A.K.E.E.V., Sharma A.K. Prevalence of hypertension and subtypes in an Indian rural population: Clinical and electrocardiographic correlates. J. Hum. Hypertens. 1994;8:823–829. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

