Nitro-fatty acids as activators of hSIRT6 deacetylase activity
- PMID: 33122195
- PMCID: PMC7939442
- DOI: 10.1074/jbc.RA120.014883
Nitro-fatty acids as activators of hSIRT6 deacetylase activity
Abstract
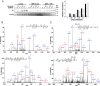
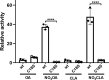
Sirtuin 6, SIRT6, is critical for both glucose and lipid homeostasis and is involved in maintaining genomic stability under conditions of oxidative DNA damage such as those observed in age-related diseases. There is an intense search for modulators of SIRT6 activity, however, not many specific activators have been reported. Long acyl-chain fatty acids have been shown to increase the weak in vitro deacetylase activity of SIRT6 but this effect is modest at best. Herein we report that electrophilic nitro-fatty acids (nitro-oleic acid and nitro-conjugated linoleic acid) potently activate SIRT6. Binding of the nitro-fatty acid to the hydrophobic crevice of the SIRT6 active site exerted a moderate activation (2-fold at 20 μm), similar to that previously reported for non-nitrated fatty acids. However, covalent Michael adduct formation with Cys-18, a residue present at the N terminus of SIRT6 but absent from other isoforms, induced a conformational change that resulted in a much stronger activation (40-fold at 20 μm). Molecular modeling of the resulting Michael adduct suggested stabilization of the co-substrate and acyl-binding loops as a possible additional mechanism of SIRT6 activation by the nitro-fatty acid. Importantly, treatment of cells with nitro-oleic acid promoted H3K9 deacetylation, whereas oleic acid had no effect. Altogether, our results show that nitrated fatty acids can be considered a valuable tool for specific SIRT6 activation, and that SIRT6 should be considered as a molecular target for in vivo actions of these anti-inflammatory nitro-lipids.
Keywords: Michael addition; SIRT6; activator; cysteine covalent modification; docking; enzyme activation; enzyme activator; enzyme kinetics; enzyme mechanism; enzyme structure; fatty acid; fatty acids; histone deacetylase (HDAC); histone deacetylation; nitro-fatty acid; nitro-fatty acids; sirtuin.
© 2020 Carreño et al.
Conflict of interest statement
Conflict of interest—The authors declare that they have no conflict of interest with the contents of this article.
Figures




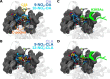


References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

