When a foreign gene meets its native counterpart: computational biophysics analysis of two PgiC loci in the grass Festuca ovina
- PMID: 33127989
- PMCID: PMC7599235
- DOI: 10.1038/s41598-020-75650-0
When a foreign gene meets its native counterpart: computational biophysics analysis of two PgiC loci in the grass Festuca ovina
Abstract
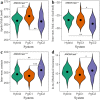
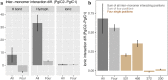
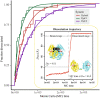
Duplicative horizontal gene transfer may bring two previously separated homologous genes together, which may raise questions about the interplay between the gene products. One such gene pair is the "native" PgiC1 and "foreign" PgiC2 in the perennial grass Festuca ovina. Both PgiC1 and PgiC2 encode cytosolic phosphoglucose isomerase, a dimeric enzyme whose proper binding is functionally essential. Here, we use biophysical simulations to explore the inter-monomer binding of the two homodimers and the heterodimer that can be produced by PgiC1 and PgiC2 in F. ovina. Using simulated native-state ensembles, we examine the structural properties and binding tightness of the dimers. In addition, we investigate their ability to withstand dissociation when pulled by a force. Our results suggest that the inter-monomer binding is tighter in the PgiC2 than the PgiC1 homodimer, which could explain the more frequent occurrence of the foreign PgiC2 homodimer in dry habitats. We further find that the PgiC1 and PgiC2 monomers are compatible with heterodimer formation; the computed binding tightness is comparable to that of the PgiC1 homodimer. Enhanced homodimer stability and capability of heterodimer formation with PgiC1 are properties of PgiC2 that may contribute to the retaining of the otherwise redundant PgiC2 gene.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
The introgression of a functional nuclear gene from Poa to Festuca ovina.Proc Biol Sci. 2006 Feb 22;273(1585):395-9. doi: 10.1098/rspb.2005.3355. Proc Biol Sci. 2006. PMID: 16615204 Free PMC article.
-
Evidence for Positive Selection within the PgiC1 Locus in the Grass Festuca ovina.PLoS One. 2015 May 6;10(5):e0125831. doi: 10.1371/journal.pone.0125831. eCollection 2015. PLoS One. 2015. PMID: 25946223 Free PMC article.
-
Origin and timing of the horizontal transfer of a PgiC gene from Poa to Festuca ovina.Mol Phylogenet Evol. 2008 Mar;46(3):890-6. doi: 10.1016/j.ympev.2007.11.031. Epub 2007 Dec 10. Mol Phylogenet Evol. 2008. PMID: 18226929
-
Contrasting patterns of nucleotide polymorphism suggest different selective regimes within different parts of the PgiC1 gene in Festuca ovina L.Hereditas. 2017 May 18;154:11. doi: 10.1186/s41065-017-0032-6. eCollection 2017. Hereditas. 2017. PMID: 28529468 Free PMC article.
-
A decade of "chromosome painting" in Lolium and Festuca.Cytogenet Genome Res. 2005;109(1-3):393-9. doi: 10.1159/000082425. Cytogenet Genome Res. 2005. PMID: 15753602 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

