A signature of 18 immune-related gene pairs to predict the prognosis of pancreatic cancer patients
- PMID: 33128857
- PMCID: PMC7654420
- DOI: 10.1002/iid3.363
A signature of 18 immune-related gene pairs to predict the prognosis of pancreatic cancer patients
Abstract
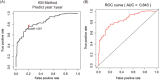
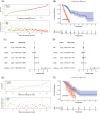
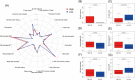
Pancreatic cancer is one of the most lethal malignancies. With the promising prospects conveyed by immunotherapy in cancers, we aimed to construct an immune-related gene pairs (IRGPs) signature to predict the prognosis of pancreatic cancer patients. We downloaded clinical and transcriptional data of pancreatic cancer patients from The Cancer Genome Atlas data set as the training group and GSE57495 data set as the verification group. We filtered immune-related transcriptional data by IMMPORT. With the assistance of lasso penalized Cox regression, we constructed our prognostic IRGPs signature and divided all samples into high-/low-risk groups by receiver operating characteristic curve for further comparisons. The comparisons between high- and low-risk groups including survival rate, multivariate, and univariate Cox proportional-hazards analysis, infiltration of immune cells, and Gene Set Enrichment Analysis (GSEA). Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) are facilitated to analyze the proceedings in which our IRGPs signature may involve in. The results revealed that 18 IRGPs were defined as our prognostic signature. The prognostic value of this IRGPs signature was verified from the GSE57495 data set. We further demonstrated the independent prognostic value of this IRGPs signature. The contents of six immune cells between high-/low-risk groups were different, which was associated with the progression of diverse cancers. Results from GO, KEGG, and GSEA revealed that this IRGPs signature was involved in extracellular space, immune response, cancer pathways, cation channel, and gated channel activities. Evidently, this IRGPs signature will provide remarkable value for the therapy of pancreatic cancer patients.
Keywords: immune-related gene pair; pancreatic cancer; prognostic signature.
© 2020 The Authors. Immunity, Inflammation and Disease published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that there are no conflict of interests.
Figures




Similar articles
-
A signature of immune-related gene pairs (IRGPs) for risk stratification and prognosis of oral cancer patients.World J Surg Oncol. 2022 Jul 8;20(1):227. doi: 10.1186/s12957-022-02630-1. World J Surg Oncol. 2022. PMID: 35804390 Free PMC article.
-
Development and Validation of a Five-immune Gene Pair Signature in Endometrial Carcinoma.Comb Chem High Throughput Screen. 2021;24(2):233-245. doi: 10.2174/1386207323999200729113641. Comb Chem High Throughput Screen. 2021. PMID: 32729416
-
Development and Verification of an Immune-Related Gene Pairs Prognostic Signature in Hepatocellular Carcinoma.Front Mol Biosci. 2021 Oct 1;8:715728. doi: 10.3389/fmolb.2021.715728. eCollection 2021. Front Mol Biosci. 2021. PMID: 34660693 Free PMC article.
-
Development and Validation of an Individualized Immune Prognostic Signature for Recurrent Prostate Cancer.Comb Chem High Throughput Screen. 2021;24(1):98-108. doi: 10.2174/1386207323666200627212820. Comb Chem High Throughput Screen. 2021. PMID: 32593277
-
Development and Validation of an Inflammatory Response-Related Gene Signature for Predicting the Prognosis of Pancreatic Adenocarcinoma.Inflammation. 2022 Aug;45(4):1732-1751. doi: 10.1007/s10753-022-01657-6. Epub 2022 Mar 23. Inflammation. 2022. PMID: 35322324
Cited by
-
An immune-related gene prognostic risk index for pancreatic adenocarcinoma.Front Immunol. 2022 Jul 26;13:945878. doi: 10.3389/fimmu.2022.945878. eCollection 2022. Front Immunol. 2022. PMID: 35958614 Free PMC article.
-
The immune-related prognostic gene AIM2 promotes pancreatic cancer progression via inflammasome.Sci Rep. 2025 Aug 13;15(1):29733. doi: 10.1038/s41598-025-12651-x. Sci Rep. 2025. PMID: 40804050 Free PMC article.
-
Construction of a new immune-related lncRNA model and prediction of treatment and survival prognosis of human colon cancer.World J Surg Oncol. 2022 Mar 6;20(1):71. doi: 10.1186/s12957-022-02508-2. World J Surg Oncol. 2022. PMID: 35249533 Free PMC article.
-
Nanoparticle-Based Therapeutic Strategies for Enhanced Pancreatic Ductal Adenocarcinoma Immunotherapy.Pharmaceutics. 2022 Sep 24;14(10):2033. doi: 10.3390/pharmaceutics14102033. Pharmaceutics. 2022. PMID: 36297467 Free PMC article. Review.
-
Systematic Analysis of Molecular Subtypes Based on the Expression Profile of Immune-Related Genes in Pancreatic Cancer.Oxid Med Cell Longev. 2022 Dec 15;2022:3124122. doi: 10.1155/2022/3124122. eCollection 2022. Oxid Med Cell Longev. 2022. PMID: 36567857 Free PMC article.
References
-
- Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016;388(10039):73‐85. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

