Piecewise All-Atom SMD Simulations Reveal Key Secondary Structures in Luciferase Unfolding Pathway
- PMID: 33130123
- PMCID: PMC7732773
- DOI: 10.1016/j.bpj.2020.10.023
Piecewise All-Atom SMD Simulations Reveal Key Secondary Structures in Luciferase Unfolding Pathway
Abstract
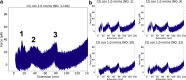
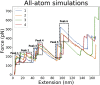
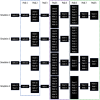
Although the folding of single-domain proteins is well characterized theoretically and experimentally, the folding of large multidomain proteins is less well known. Firefly luciferase, a 550 residue three-domain protein, has been commonly used as a substrate to study chaperone reactions and as a model system for the study of folding of long polypeptide chains, including related phenomena such as cotranslational folding. Despite being characterized by various experimental techniques, the atomic-level contributions of various secondary structures of luciferase to its fold's mechanical stability remain unknown. Here, we developed a piecewise approach for all-atom steered molecular dynamics simulations to examine specific secondary structures that resist mechanical unfolding while minimizing the amount of computational resources required by the large water box of standard all-atom steered molecular dynamics simulations. We validated the robustness of this approach with a small NI3C protein and used our approach to elucidate the specific secondary structures that provide the largest contributions to luciferase mechanostability. In doing so, we show that piecewise all-atom steered molecular dynamics simulations can provide novel atomic resolution details regarding mechanostability and can serve as a platform for novel mutagenesis studies as well as a point for comparison with high-resolution force spectroscopy experiments.
Copyright © 2020 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures




Similar articles
-
Chaperones rescue luciferase folding by separating its domains.J Biol Chem. 2014 Oct 10;289(41):28607-18. doi: 10.1074/jbc.M114.582049. Epub 2014 Aug 26. J Biol Chem. 2014. PMID: 25160632 Free PMC article.
-
Reconstruction of mechanical unfolding and refolding pathways of proteins with atomic force spectroscopy and computer simulations.Methods. 2022 Jan;197:39-53. doi: 10.1016/j.ymeth.2021.05.012. Epub 2021 May 18. Methods. 2022. PMID: 34020035
-
Competing Pathways and Multiple Folding Nuclei in a Large Multidomain Protein, Luciferase.Biophys J. 2017 May 9;112(9):1829-1840. doi: 10.1016/j.bpj.2017.03.028. Biophys J. 2017. PMID: 28494954 Free PMC article.
-
Successes and challenges in simulating the folding of large proteins.J Biol Chem. 2020 Jan 3;295(1):15-33. doi: 10.1074/jbc.REV119.006794. Epub 2019 Nov 11. J Biol Chem. 2020. PMID: 31712314 Free PMC article. Review.
-
Dynamics of protein folding: probing the kinetic network of folding-unfolding transitions with experiment and theory.Biochim Biophys Acta. 2011 Aug;1814(8):1001-20. doi: 10.1016/j.bbapap.2010.09.013. Epub 2010 Sep 29. Biochim Biophys Acta. 2011. PMID: 20883829 Review.
Cited by
-
Implementation of Telescoping Boxes in Adaptive Steered Molecular Dynamics.J Chem Theory Comput. 2022 Aug 9;18(8):4649-4659. doi: 10.1021/acs.jctc.2c00498. Epub 2022 Jul 13. J Chem Theory Comput. 2022. PMID: 35830368 Free PMC article.
-
Tandem repeats of highly bioluminescent NanoLuc are refolded noncanonically by the Hsp70 machinery.Protein Sci. 2024 Feb;33(2):e4895. doi: 10.1002/pro.4895. Protein Sci. 2024. PMID: 38284490 Free PMC article.
-
Molecular Mechanism of Interaction between DNA Aptamer and Receptor-Binding Domain of Severe Acute Respiratory Syndrome Coronavirus 2 Variants Revealed by Steered Molecular Dynamics Simulations.Molecules. 2024 May 9;29(10):2215. doi: 10.3390/molecules29102215. Molecules. 2024. PMID: 38792076 Free PMC article.
References
-
- Dobson C.M. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Dill K.A., MacCallum J.L. The protein-folding problem, 50 years on. Science. 2012;338:1042–1046. - PubMed
-
- Anfinsen C.B. Principles that govern the folding of protein chains. Science. 1973;181:223–230. - PubMed
-
- Onuchic J.N., Luthey-Schulten Z., Wolynes P.G. Theory of protein folding: the energy landscape perspective. Annu. Rev. Phys. Chem. 1997;48:545–600. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

