Structure of S. pombe telomerase protein Pof8 C-terminal domain is an xRRM conserved among LARP7 proteins
- PMID: 33131423
- PMCID: PMC8244769
- DOI: 10.1080/15476286.2020.1836891
Structure of S. pombe telomerase protein Pof8 C-terminal domain is an xRRM conserved among LARP7 proteins
Abstract
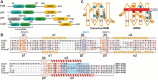
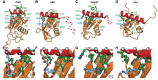
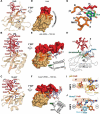
La-related proteins 7 (LARP7) are a class of RNA chaperones that bind the 3' ends of RNA and are constitutively associated with their specific target RNAs. In metazoa, Larp7 binds to the long non-coding 7SK RNA as a core component of the 7SK RNP, a major regulator of eukaryotic transcription. In the ciliate Tetrahymena the LARP7 protein p65 is a component of telomerase, an essential ribonucleoprotein complex that maintains the telomeric DNA at eukaryotic chromosome ends. p65 is important for the ordered assembly of telomerase RNA (TER) with telomerase reverse transcriptase. Unexpectedly, Schizosaccharomyces pombe Pof8 was recently identified as a LARP7 protein and a core component of fission yeast telomerase essential for biogenesis. LARP7 proteins have a conserved N-terminal La motif and RRM1 (La module) and C-terminal RRM2 with specific RNA substrate recognition attributed to RRM2, first structurally characterized in p65 as an atypical RRM named xRRM. Here we present the X-ray crystal structure and NMR studies of S. pombe Pof8 RRM2. Sequence and structure comparison of Pof8 RRM2 to p65 and human Larp7 xRRMs reveals conserved features for RNA binding with the main variability in the length of the non-canonical helix α3. This study shows that Pof8 has conserved xRRM features, providing insight into TER recognition and the defining characteristics of the xRRM.
Keywords: 7SK; LARP; La protein; NMR; RNA; RRM; X-ray crystallography; telomerase; xRRM.
Figures


Similar articles
-
LARP7-like protein Pof8 regulates telomerase assembly and poly(A)+TERRA expression in fission yeast.Nat Commun. 2018 Feb 8;9(1):586. doi: 10.1038/s41467-018-02874-0. Nat Commun. 2018. PMID: 29422503 Free PMC article.
-
xRRM: a new class of RRM found in the telomerase La family protein p65.RNA Biol. 2013 Mar;10(3):353-9. doi: 10.4161/rna.23608. Epub 2013 Jan 17. RNA Biol. 2013. PMID: 23328630 Free PMC article. Review.
-
Structural basis for telomerase RNA recognition and RNP assembly by the holoenzyme La family protein p65.Mol Cell. 2012 Jul 13;47(1):16-26. doi: 10.1016/j.molcel.2012.05.018. Epub 2012 Jun 14. Mol Cell. 2012. PMID: 22705372 Free PMC article.
-
Pof8 is a La-related protein and a constitutive component of telomerase in fission yeast.Nat Commun. 2018 Feb 8;9(1):587. doi: 10.1038/s41467-017-02284-8. Nat Commun. 2018. PMID: 29422664 Free PMC article.
-
Stabilize and connect: the role of LARP7 in nuclear non-coding RNA metabolism.RNA Biol. 2021 Feb;18(2):290-303. doi: 10.1080/15476286.2020.1767952. Epub 2020 Jun 3. RNA Biol. 2021. PMID: 32401147 Free PMC article. Review.
Cited by
-
Progress in 7SK ribonucleoprotein structural biology.Front Mol Biosci. 2023 Mar 27;10:1154622. doi: 10.3389/fmolb.2023.1154622. eCollection 2023. Front Mol Biosci. 2023. PMID: 37051324 Free PMC article. Review.
-
The fission yeast methyl phosphate capping enzyme Bmc1 guides 2'-O-methylation of the U6 snRNA.Nucleic Acids Res. 2023 Sep 8;51(16):8805-8819. doi: 10.1093/nar/gkad563. Nucleic Acids Res. 2023. PMID: 37403782 Free PMC article.
-
The methyl phosphate capping enzyme Bmc1/Bin3 is a stable component of the fission yeast telomerase holoenzyme.Nat Commun. 2022 Mar 11;13(1):1277. doi: 10.1038/s41467-022-28985-3. Nat Commun. 2022. PMID: 35277511 Free PMC article.
-
LARP1 and LARP4: up close with PABP for mRNA 3' poly(A) protection and stabilization.RNA Biol. 2021 Feb;18(2):259-274. doi: 10.1080/15476286.2020.1868753. Epub 2021 Jan 31. RNA Biol. 2021. PMID: 33522422 Free PMC article. Review.
-
Arabidopsis retains vertebrate-type telomerase accessory proteins via a plant-specific assembly.Nucleic Acids Res. 2021 Sep 20;49(16):9496-9507. doi: 10.1093/nar/gkab699. Nucleic Acids Res. 2021. PMID: 34403479 Free PMC article.
References
-
- Stefano JE. Purified lupus antigen La recognizes an oligouridylate stretch common to the 3ʹ termini of RNA polymerase III transcripts. Cell. 1984;36:145–154. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
