The Interrelationship Between Microbiota and Peptides During Ripening as a Driver for Parmigiano Reggiano Cheese Quality
- PMID: 33133050
- PMCID: PMC7561718
- DOI: 10.3389/fmicb.2020.581658
The Interrelationship Between Microbiota and Peptides During Ripening as a Driver for Parmigiano Reggiano Cheese Quality
Abstract
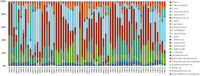
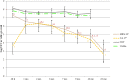
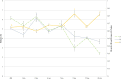
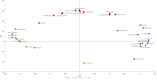
Cheese microbiota contribute significantly to the final characteristics of cheeses due to the growth and interaction between cheese microorganisms during processing and ripening. For raw milk cheeses, such as Parmigiano Reggiano (PR), the microbiota derive from the raw milk itself, the dairy environment, and the starter. The process of cheese making and time of ripening shape this complex ecosystem through the selection of different species and biotypes that will drive the quality of the final product by performing functions of their metabolism such as proteolysis. The diversity in the final peptide and amino acid composition of the cheese is thus mostly linked to the diversity of this microbiota. The purpose of this study was to get more insight into the factors affecting PR cheese diversity and, more specifically, to evaluate whether the composition of the bacterial community of cheeses along with the specific peptide composition are more affected by the ripening times or by the cheese making process. To this end, the microbiota and the peptide fractions of 69 cheese samples (from curd to cheese ripened 24 months) were analyzed during 6 complete PR production cycles, which were performed in six different dairies located in the PR production area. The relation among microbial dynamics, peptide evolution, and ripening times were investigated in this unique and tightly controlled production and sampling set up. The study of microbial and peptide moieties in products from different dairies - from curd to at least 12 months, the earliest time from which the cheese can be sold, and up to a maximum of 24 months of ripening - highlighted the presence of differences between samples coming from different dairies, probably due to small differences in the cheese making process. Besides these differences, however, ripening time had by far the greatest impact on microbial dynamics and, consequently, on peptide composition.
Keywords: Parmigiano Reggiano; cheese microbiota; cheese peptides; cheese ripening; raw milk cheese.
Copyright © 2020 Bottari, Levante, Bancalari, Sforza, Bottesini, Prandi, De Filippis, Ercolini, Nocetti and Gatti.
Figures




Similar articles
-
Peptide profile of Parmigiano Reggiano cheese after simulated gastrointestinal digestion: From quality drivers to functional compounds.Front Microbiol. 2022 Aug 23;13:966239. doi: 10.3389/fmicb.2022.966239. eCollection 2022. Front Microbiol. 2022. PMID: 36081785 Free PMC article.
-
Cultivable microbial diversity, peptide profiles, and bio-functional properties in Parmigiano Reggiano cheese.Front Microbiol. 2024 Mar 19;15:1342180. doi: 10.3389/fmicb.2024.1342180. eCollection 2024. Front Microbiol. 2024. PMID: 38567075 Free PMC article.
-
Cheese peptidomics: a detailed study on the evolution of the oligopeptide fraction in Parmigiano-Reggiano cheese from curd to 24 months of aging.J Dairy Sci. 2012 Jul;95(7):3514-26. doi: 10.3168/jds.2011-5046. J Dairy Sci. 2012. PMID: 22720910
-
Invited review: Microbial evolution in raw-milk, long-ripened cheeses produced using undefined natural whey starters.J Dairy Sci. 2014 Feb;97(2):573-91. doi: 10.3168/jds.2013-7187. Epub 2013 Dec 2. J Dairy Sci. 2014. PMID: 24290824 Review.
-
The Microfloras and Sensory Profiles of Selected Protected Designation of Origin Italian Cheeses.Microbiol Spectr. 2014 Feb;2(1):CM-0007-2012. doi: 10.1128/microbiolspec.CM-0007-2012. Microbiol Spectr. 2014. PMID: 26082116 Review.
Cited by
-
Genome Sequencing of five Lacticaseibacillus Strains and Analysis of Type I and II Toxin-Antitoxin System Distribution.Microorganisms. 2021 Mar 21;9(3):648. doi: 10.3390/microorganisms9030648. Microorganisms. 2021. PMID: 33800997 Free PMC article.
-
Lactic acid bacteria in cow raw milk for cheese production: Which and how many?Front Microbiol. 2023 Jan 12;13:1092224. doi: 10.3389/fmicb.2022.1092224. eCollection 2022. Front Microbiol. 2023. PMID: 36713157 Free PMC article. Review.
-
Microbiome profiling of Grana Padano and Parmigiano Reggiano cheeses reveals cheese-specific biomarkers, psychobiotic potential, and bioprotective activities.NPJ Biofilms Microbiomes. 2025 Aug 27;11(1):177. doi: 10.1038/s41522-025-00815-6. NPJ Biofilms Microbiomes. 2025. PMID: 40858622 Free PMC article.
-
Peptide profile of Parmigiano Reggiano cheese after simulated gastrointestinal digestion: From quality drivers to functional compounds.Front Microbiol. 2022 Aug 23;13:966239. doi: 10.3389/fmicb.2022.966239. eCollection 2022. Front Microbiol. 2022. PMID: 36081785 Free PMC article.
-
Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese.Foods. 2023 Oct 16;12(20):3796. doi: 10.3390/foods12203796. Foods. 2023. PMID: 37893689 Free PMC article.
References
-
- Alessandria V., Ferrocino I., De Filippis F., Fontana M., Rantsiou K., Ercolini D., et al. (2016). Microbiota of an Italian Grana-like cheese during manufacture and ripening, unraveled by 16S rRNA-based approaches. Appl. Environ. Microbiol. 82 3988–3995. 10.1128/AEM.00999-16.Editor - DOI - PMC - PubMed
-
- Berni Canani R., De Filippis F., Nocerino R., Laiola M., Paparo L., Calignano A., et al. (2017). Specific signatures of the gut microbiota and increased levels of butyrate in children treated with fermented cow’s milk containing heat-killed Lactobacillus paracasei CBA L74. App. Environ. Microbiol. 83 e1206–e1217. 10.1128/AEM.01206-17 - DOI - PMC - PubMed
-
- Bertani G., Levante A., Lazzi C., Bottari B., Gatti M., Neviani E. (2020). Dynamics of a natural bacterial community under technological and environmental pressures: the case of natural whey starter for parmigiano reggiano cheese. Food Res. Int. 129:108860. 10.1016/j.foodres.2019.108860 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

