Small hydrophobic viral proteins involved in intercellular movement of diverse plant virus genomes
- PMID: 33134746
- PMCID: PMC7595835
- DOI: 10.3934/microbiol.2020019
Small hydrophobic viral proteins involved in intercellular movement of diverse plant virus genomes
Abstract
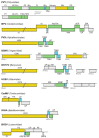
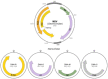
Most plant viruses code for movement proteins (MPs) targeting plasmodesmata to enable cell-to-cell and systemic spread in infected plants. Small membrane-embedded MPs have been first identified in two viral transport gene modules, triple gene block (TGB) coding for an RNA-binding helicase TGB1 and two small hydrophobic proteins TGB2 and TGB3 and double gene block (DGB) encoding two small polypeptides representing an RNA-binding protein and a membrane protein. These findings indicated that movement gene modules composed of two or more cistrons may encode the nucleic acid-binding protein and at least one membrane-bound movement protein. The same rule was revealed for small DNA-containing plant viruses, namely, viruses belonging to genus Mastrevirus (family Geminiviridae) and the family Nanoviridae. In multi-component transport modules the nucleic acid-binding MP can be viral capsid protein(s), as in RNA-containing viruses of the families Closteroviridae and Potyviridae. However, membrane proteins are always found among MPs of these multicomponent viral transport systems. Moreover, it was found that small membrane MPs encoded by many viruses can be involved in coupling viral replication and cell-to-cell movement. Currently, the studies of evolutionary origin and functioning of small membrane MPs is regarded as an important pre-requisite for understanding of the evolution of the existing plant virus transport systems. This paper represents the first comprehensive review which describes the whole diversity of small membrane MPs and presents the current views on their role in plant virus movement.
Keywords: DNA genome; RNA genome; evolution; hydrophobic protein; intercellular movement; plant virus; protein motifs; trans-membrane domain; triple gene block.
© 2020 the Author(s), licensee AIMS Press.
Conflict of interest statement
Conflict of Interest: The authors declare that they have no conflict of interest.
Figures
Similar articles
-
Phylogenetic relationship of some "accessory" helicases of plant positive-stranded RNA viruses: toward understanding the evolution of triple gene block.Front Microbiol. 2015 May 19;6:508. doi: 10.3389/fmicb.2015.00508. eCollection 2015. Front Microbiol. 2015. PMID: 26042118 Free PMC article.
-
Non-replicative Integral Membrane Proteins Encoded by Plant Alpha-Like Viruses: Emergence of Diverse Orphan ORFs and Movement Protein Genes.Front Plant Sci. 2017 Oct 27;8:1820. doi: 10.3389/fpls.2017.01820. eCollection 2017. Front Plant Sci. 2017. PMID: 29163564 Free PMC article.
-
Similarities in intracellular transport of plant viral movement proteins BMB2 and TGB3.J Gen Virol. 2017 Sep;98(9):2379-2391. doi: 10.1099/jgv.0.000914. Epub 2017 Sep 4. J Gen Virol. 2017. PMID: 28869000
-
Role of plant virus movement proteins.Methods Mol Biol. 2008;451:33-54. doi: 10.1007/978-1-59745-102-4_3. Methods Mol Biol. 2008. PMID: 18370246 Review.
-
Cell-to-cell movement of plant viruses. Insights from amino acid sequence comparisons of movement proteins and from analogies with cellular transport systems.Arch Virol. 1993;133(3-4):239-57. doi: 10.1007/BF01313766. Arch Virol. 1993. PMID: 8257287 Free PMC article. Review.
Cited by
-
Alphaflexiviridae in Focus: Genomic Signatures, Conserved Elements and Viral-Driven Cellular Remodeling.Viruses. 2025 Apr 24;17(5):611. doi: 10.3390/v17050611. Viruses. 2025. PMID: 40431623 Free PMC article. Review.
-
Complete sequence and genome characterization of miscanthus virus M, a new betaflexivirus from Miscanthus sp.Arch Virol. 2024 Jan 12;169(2):27. doi: 10.1007/s00705-024-05966-z. Arch Virol. 2024. PMID: 38214767
-
Uncovering Plant Virus Species Forming Novel Provisional Taxonomic Units Related to the Family Benyviridae.Viruses. 2022 Nov 29;14(12):2680. doi: 10.3390/v14122680. Viruses. 2022. PMID: 36560684 Free PMC article.
-
A Genetic Study of Spillovers in the Bean Common Mosaic Subgroup of Potyviruses.Viruses. 2024 Aug 23;16(9):1351. doi: 10.3390/v16091351. Viruses. 2024. PMID: 39339828 Free PMC article.
-
Viruses Infecting Greenhood Orchids (Pterostylidinae) in Eastern Australia.Viruses. 2022 Feb 10;14(2):365. doi: 10.3390/v14020365. Viruses. 2022. PMID: 35215958 Free PMC article.
References
-
- Tilsner J, Nicolas W, Rosado A, et al. Staying tight: plasmodesmal membrane contact sites and the control of cell-to-cell connectivity in plants. Annu Rev Plant Biol. 2016;67:337–364. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
