Antiviral and Immunomodulatory Effects of Phytochemicals from Honey against COVID-19: Potential Mechanisms of Action and Future Directions
- PMID: 33138197
- PMCID: PMC7672575
- DOI: 10.3390/molecules25215017
Antiviral and Immunomodulatory Effects of Phytochemicals from Honey against COVID-19: Potential Mechanisms of Action and Future Directions
Abstract
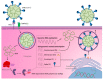
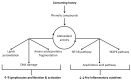
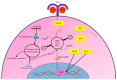
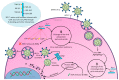
The new coronavirus disease (COVID-19), caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), has recently put the world under stress, resulting in a global pandemic. Currently, there are no approved treatments or vaccines, and this severe respiratory illness has cost many lives. Despite the established antimicrobial and immune-boosting potency described for honey, to date there is still a lack of evidence about its potential role amid COVID-19 outbreak. Based on the previously explored antiviral effects and phytochemical components of honey, we review here evidence for its role as a potentially effective natural product against COVID-19. Although some bioactive compounds in honey have shown potential antiviral effects (i.e., methylglyoxal, chrysin, caffeic acid, galangin and hesperidinin) or enhancing antiviral immune responses (i.e., levan and ascorbic acid), the mechanisms of action for these compounds are still ambiguous. To the best of our knowledge, this is the first work exclusively summarizing all these bioactive compounds with their probable mechanisms of action as antiviral agents, specifically against SARS-CoV-2.
Keywords: 2019-nCoV; SARS-CoV-2; antiviral activity; antiviral agent; antiviral immunity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- WHO Timeline—COVID-19. [(accessed on 7 June 2020)]; Available online: https://www.who.int/news-room/detail/27-04-2020-who-timeline---covid-19.
-
- Coronavirus Disease (COVID-2019) Situation Reports. [(accessed on 14 September 2020)]; Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situatio....
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

