Necessity for detection of SARS-CoV-2 RNA in multiple types of specimens for the discharge of the patients with COVID-19
- PMID: 33138834
- PMCID: PMC7605325
- DOI: 10.1186/s12967-020-02580-w
Necessity for detection of SARS-CoV-2 RNA in multiple types of specimens for the discharge of the patients with COVID-19
Abstract
Background: The SARS-CoV-2 RNA was detected positive again after discharged from hospital in some COVID-19 patients, with or without clinical symptoms such as fever or dry cough.
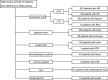
Methods: 1008 severe COVID-19 patients, with SARS-CoV-2 RNA positive detected with the mixed specimen of nasopharyngeal swab and oropharyngeal swab by real-time fluorescence quantitative PCR (RT-qPCR), were selected to monitor SARS-CoV-2 RNA with the 12 types of specimens by RT-qPCR during hospitalization. All of 20 discharged cases with COVID-19 were selected to detect SARS-CoV-2 RNA in isolation period with 7 types of specimens by RT-qPCR before releasing the isolation period.
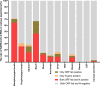
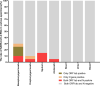
Results: Of the enrolled 1008 severe patients, the nasopharyngeal swab specimens showed the highest positive rate of SARS-CoV-2 RNA (71.06%), followed by alveolar lavage fluid (66.67%), oropharyngeal swab (30.77%), sputum (28.53%), urine (16.30%), blood (12.5%), stool (12.21%), anal swab (11.22%) and corneal secretion (2.99%), and SARS-CoV-2 RNA couldn't be detected in other types of specimen in this study. Of the 20 discharged cases during the isolation period, the positive rate of SARS-CoV-2 RNA was 30% (6/20): 2 cases were positive in sputum at the eighth and ninth day after discharge, respectively, 1 case was positive in nasopharynx swab at the sixth day after discharge, 1 case was positive in anal swab at the eighth day after discharge, and 1 case was positive in 3 specimens (nasopharynx swab, oropharynx swab and sputum) simultaneously at the fourth day after discharge, and no positive SARS-CoV-2 RNA was detected in other specimens including stool, urine and blood at the discharged patients.
Conclusions: SARS-CoV-2 RNA should be detected in multiple specimens, such as nasopharynx swab, oropharynx swab, sputum, and if necessary, stool and anal swab specimens should be performed simultaneously at discharge when the patients were considered for clinical cure and before releasing the isolation period.
Keywords: COVID-19; Discharge criteria; Multiple specimens; RT-qPCR; SARS-CoV-2.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Value of swab types and collection time on SARS-COV-2 detection using RT-PCR assay.J Virol Methods. 2020 Dec;286:113974. doi: 10.1016/j.jviromet.2020.113974. Epub 2020 Sep 16. J Virol Methods. 2020. PMID: 32949663 Free PMC article.
-
Presence of viral RNA of SARS-CoV-2 in conjunctival swab specimens of COVID-19 patients.Indian J Ophthalmol. 2020 Jun;68(6):1015-1017. doi: 10.4103/ijo.IJO_1287_20. Indian J Ophthalmol. 2020. PMID: 32461418 Free PMC article.
-
Detection of SARS-CoV-2 RNA by direct RT-qPCR on nasopharyngeal specimens without extraction of viral RNA.PLoS One. 2020 Jul 24;15(7):e0236564. doi: 10.1371/journal.pone.0236564. eCollection 2020. PLoS One. 2020. PMID: 32706827 Free PMC article.
-
SARS-CoV-2 detection in different respiratory sites: A systematic review and meta-analysis.EBioMedicine. 2020 Sep;59:102903. doi: 10.1016/j.ebiom.2020.102903. Epub 2020 Jul 24. EBioMedicine. 2020. PMID: 32718896 Free PMC article.
-
[SARS-CoV-2 and Microbiological Diagnostic Dynamics in COVID-19 Pandemic].Mikrobiyol Bul. 2020 Jul;54(3):497-509. doi: 10.5578/mb.69839. Mikrobiyol Bul. 2020. PMID: 32755524 Review. Turkish.
Cited by
-
SARS-CoV-2 laboratory surveillance during the first year of the COVID-19 pandemic in southern Brazil.Rev Soc Bras Med Trop. 2023 Jan 23;56:e0146-2022. doi: 10.1590/0037-8682-0146-2022. eCollection 2023. Rev Soc Bras Med Trop. 2023. PMID: 36700597 Free PMC article.
-
Screening and confirmation tests for SARS-CoV-2: benefits and drawbacks.Beni Suef Univ J Basic Appl Sci. 2023;12(1):6. doi: 10.1186/s43088-023-00342-3. Epub 2023 Jan 11. Beni Suef Univ J Basic Appl Sci. 2023. PMID: 36647397 Free PMC article. Review.
-
Viral RNA Load and Infectivity of SARS-CoV-2 in Paired Respiratory and Oral Specimens from Symptomatic, Asymptomatic, or Postsymptomatic Individuals.Microbiol Spectr. 2022 Jun 29;10(3):e0226421. doi: 10.1128/spectrum.02264-21. Epub 2022 May 16. Microbiol Spectr. 2022. PMID: 35575498 Free PMC article.
-
Monitoring the presence and persistence of SARS-CoV-2 in water-food-environmental compartments: State of the knowledge and research needs.Environ Res. 2021 Sep;200:111373. doi: 10.1016/j.envres.2021.111373. Epub 2021 May 24. Environ Res. 2021. PMID: 34033834 Free PMC article. Review.
-
Recent Chronology of COVID-19 Pandemic.Front Public Health. 2022 May 4;10:778037. doi: 10.3389/fpubh.2022.778037. eCollection 2022. Front Public Health. 2022. PMID: 35602161 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

