Disentangling the Roles of RIM and Munc13 in Synaptic Vesicle Localization and Neurotransmission
- PMID: 33139401
- PMCID: PMC7724145
- DOI: 10.1523/JNEUROSCI.1922-20.2020
Disentangling the Roles of RIM and Munc13 in Synaptic Vesicle Localization and Neurotransmission
Abstract
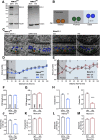
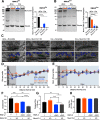
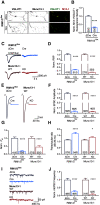
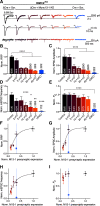
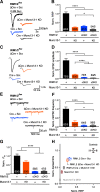
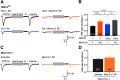
Efficient neurotransmitter release at the presynaptic terminal requires docking of synaptic vesicles to the active zone membrane and formation of fusion-competent synaptic vesicles near voltage-gated Ca2+ channels. Rab3-interacting molecule (RIM) is a critical active zone organizer, as it recruits Ca2+ channels and activates synaptic vesicle docking and priming via Munc13-1. However, our knowledge about Munc13-independent contributions of RIM to active zone functions is limited. To identify the functions that are solely mediated by RIM, we used genetic manipulations to control RIM and Munc13-1 activity in cultured hippocampal neurons from mice of either sex and compared synaptic ultrastructure and neurotransmission. We found that RIM modulates synaptic vesicle localization in the proximity of the active zone membrane independent of Munc13-1. In another step, both RIM and Munc13 mediate synaptic vesicle docking and priming. In addition, while the activity of both RIM and Munc13-1 is required for Ca2+-evoked release, RIM uniquely controls neurotransmitter release efficiency. However, activity-dependent augmentation of synaptic vesicle pool size relies exclusively on the action of Munc13s. Based on our results, we extend previous findings and propose a refined model in which RIM and Munc13-1 act in overlapping and independent stages of synaptic vesicle localization and release.SIGNIFICANCE STATEMENT The presynaptic active zone is composed of scaffolding proteins that functionally interact to localize synaptic vesicles to release sites, ensuring neurotransmission. Our current knowledge of the presynaptic active zone function relies on structure-function analysis, which has provided detailed information on the network of interactions and the impact of active zone proteins. Yet, the hierarchical, redundant, or independent cooperation of each active zone protein to synapse functions is not fully understood. Rab3-interacting molecule and Munc13 are the two key functionally interacting active zone proteins. Here, we dissected the distinct actions of Rab3-interacting molecule and Munc13-1 from both ultrastructural and physiological aspects. Our findings provide a more detailed view of how these two presynaptic proteins orchestrate their functions to achieve synaptic transmission.
Keywords: Munc13; RIM; active zone; electron microscopy; synaptic transmission; synaptic vesicle.
Copyright © 2020 Zarebidaki et al.
Figures

Similar articles
-
A Trio of Active Zone Proteins Comprised of RIM-BPs, RIMs, and Munc13s Governs Neurotransmitter Release.Cell Rep. 2020 Aug 4;32(5):107960. doi: 10.1016/j.celrep.2020.107960. Cell Rep. 2020. PMID: 32755572
-
Heterodimerization of Munc13 C2A domain with RIM regulates synaptic vesicle docking and priming.Nat Commun. 2017 May 10;8:15293. doi: 10.1038/ncomms15293. Nat Commun. 2017. PMID: 28489077 Free PMC article.
-
Munc13 supports fusogenicity of non-docked vesicles at synapses with disrupted active zones.Elife. 2022 Nov 18;11:e79077. doi: 10.7554/eLife.79077. Elife. 2022. PMID: 36398873 Free PMC article.
-
Neurotransmitter release: the last millisecond in the life of a synaptic vesicle.Neuron. 2013 Oct 30;80(3):675-90. doi: 10.1016/j.neuron.2013.10.022. Neuron. 2013. PMID: 24183019 Free PMC article. Review.
-
The presynaptic active zone.Neuron. 2012 Jul 12;75(1):11-25. doi: 10.1016/j.neuron.2012.06.012. Neuron. 2012. PMID: 22794257 Free PMC article. Review.
Cited by
-
Mechanisms of Synaptic Vesicle Exo- and Endocytosis.Biomedicines. 2022 Jul 4;10(7):1593. doi: 10.3390/biomedicines10071593. Biomedicines. 2022. PMID: 35884898 Free PMC article. Review.
-
Immune-metabolic mechanisms of post-traumatic stress disorder and atherosclerosis.Front Physiol. 2023 Feb 8;14:1123692. doi: 10.3389/fphys.2023.1123692. eCollection 2023. Front Physiol. 2023. PMID: 36846337 Free PMC article. Review.
-
The Role of Rab Proteins in Parkinson's Disease Synaptopathy.Biomedicines. 2022 Aug 10;10(8):1941. doi: 10.3390/biomedicines10081941. Biomedicines. 2022. PMID: 36009486 Free PMC article. Review.
-
RIM and MUNC13 membrane-binding domains are essential for neuropeptide secretion.J Cell Biol. 2025 Jul 7;224(7):e202409196. doi: 10.1083/jcb.202409196. Epub 2025 May 12. J Cell Biol. 2025. PMID: 40353777 Free PMC article.
-
PXF-1 promotes synapse development at the neuromuscular junction in Caenorhabditis elegans.Front Mol Neurosci. 2022 Oct 13;15:945680. doi: 10.3389/fnmol.2022.945680. eCollection 2022. Front Mol Neurosci. 2022. PMID: 36311020 Free PMC article.
References
-
- Arancillo M, Min SW, Gerber S, Münster-Wandowski A, Wu YJ, Herman M, Trimbuch T, Rah JC, Ahnert-Hilger G, Riedel D, Südhof TC, Rosenmund C (2013) Titration of Syntaxin1 in mammalian synapses reveals multiple roles in vesicle docking, priming, and release probability. J Neurosci 33:16698–16714. 10.1523/JNEUROSCI.0187-13.2013 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
