A biomimetic five-module chimeric antigen receptor (5MCAR) designed to target and eliminate antigen-specific T cells
- PMID: 33139567
- PMCID: PMC7682351
- DOI: 10.1073/pnas.2012495117
A biomimetic five-module chimeric antigen receptor (5MCAR) designed to target and eliminate antigen-specific T cells
Abstract
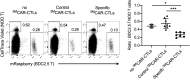
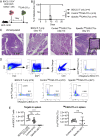
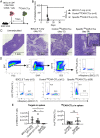
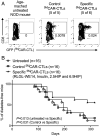
T cells express clonotypic T cell receptors (TCRs) that recognize peptide antigens in the context of class I or II MHC molecules (pMHCI/II). These receptor modules associate with three signaling modules (CD3γε, δε, and ζζ) and work in concert with a coreceptor module (either CD8 or CD4) to drive T cell activation in response to pMHCI/II. Here, we describe a first-generation biomimetic five-module chimeric antigen receptor (5MCAR). We show that 1) chimeric receptor modules built with the ectodomains of pMHCII assemble with CD3 signaling modules into complexes that redirect cytotoxic T lymphocyte (CTL) specificity and function in response to the clonotypic TCRs of pMHCII-specific CD4+ T cells, and 2) surrogate coreceptor modules enhance the function of these complexes. Furthermore, we demonstrate that adoptively transferred 5MCAR-CTLs can mitigate type I diabetes by targeting autoimmune CD4+ T cells in NOD mice. This work provides a framework for the construction of biomimetic 5MCARs that can be used as tools to study the impact of particular antigen-specific T cells in immune responses, and may hold potential for ameliorating diseases mediated by pathogenic T cells.
Keywords: 5M-CAR; CAR; T1D; TCR; pMHC.
Conflict of interest statement
Competing interest statement: M.S.K. and T.S. have disclosed an outside interest in Module Therapeutics to the University of Arizona and the Joslin Diabetes Center. Conflicts of interest resulting from this interest are being managed by The University of Arizona and Joslin Diabetes Center in accordance with their policies. M.S.K. and T.S. are inventors on patent filings covering the intellectual property tested in this study.
Figures



Similar articles
-
Chimeric antigen receptor (CAR) T cells targeting a pathogenic MHC class II:peptide complex modulate the progression of autoimmune diabetes.J Autoimmun. 2019 Jan;96:50-58. doi: 10.1016/j.jaut.2018.08.004. Epub 2018 Aug 16. J Autoimmun. 2019. PMID: 30122420 Free PMC article.
-
Enhanced antilymphoma efficacy of CD19-redirected influenza MP1-specific CTLs by cotransfer of T cells modified to present influenza MP1.Blood. 2005 Feb 15;105(4):1622-31. doi: 10.1182/blood-2004-03-1208. Epub 2004 Oct 26. Blood. 2005. PMID: 15507526
-
Different T cell receptor affinity thresholds and CD8 coreceptor dependence govern cytotoxic T lymphocyte activation and tetramer binding properties.J Biol Chem. 2007 Aug 17;282(33):23799-810. doi: 10.1074/jbc.M700976200. Epub 2007 May 31. J Biol Chem. 2007. PMID: 17540778
-
Signaling from T cell receptors (TCRs) and chimeric antigen receptors (CARs) on T cells.Cell Mol Immunol. 2020 Jun;17(6):600-612. doi: 10.1038/s41423-020-0470-3. Epub 2020 May 25. Cell Mol Immunol. 2020. PMID: 32451454 Free PMC article. Review.
-
Interplay between the TCR/CD3 complex and CD4 or CD8 in the activation of cytotoxic T lymphocytes.Immunol Rev. 1989 Jun;109:119-41. doi: 10.1111/j.1600-065x.1989.tb00022.x. Immunol Rev. 1989. PMID: 2527803 Review.
Cited by
-
Multipurposing CARs: Same engine, different vehicles.Mol Ther. 2022 Apr 6;30(4):1381-1395. doi: 10.1016/j.ymthe.2022.02.012. Epub 2022 Feb 11. Mol Ther. 2022. PMID: 35151842 Free PMC article. Review.
-
Diverse potential of chimeric antigen receptor-engineered cell therapy: Beyond cancer.Clin Transl Med. 2025 Apr;15(4):e70306. doi: 10.1002/ctm2.70306. Clin Transl Med. 2025. PMID: 40205818 Free PMC article. Review.
-
The CD4 transmembrane GGXXG and juxtamembrane (C/F)CV+C motifs mediate pMHCII-specific signaling independently of CD4-LCK interactions.bioRxiv [Preprint]. 2023 Nov 29:2023.05.05.539613. doi: 10.1101/2023.05.05.539613. bioRxiv. 2023. Update in: Elife. 2024 Apr 19;12:RP88225. doi: 10.7554/eLife.88225. PMID: 37214965 Free PMC article. Updated. Preprint.
-
Cell-based therapies for rheumatoid arthritis: opportunities and challenges.Ther Adv Musculoskelet Dis. 2022 May 23;14:1759720X221100294. doi: 10.1177/1759720X221100294. eCollection 2022. Ther Adv Musculoskelet Dis. 2022. PMID: 35634355 Free PMC article. Review.
-
Current progress in CAR-based therapy for kidney disease.Front Immunol. 2024 Aug 20;15:1408718. doi: 10.3389/fimmu.2024.1408718. eCollection 2024. Front Immunol. 2024. PMID: 39234257 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

