Modelling co-translational dimerization for programmable nonlinearity in synthetic biology
- PMID: 33143595
- PMCID: PMC7729047
- DOI: 10.1098/rsif.2020.0561
Modelling co-translational dimerization for programmable nonlinearity in synthetic biology
Abstract
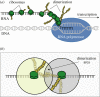
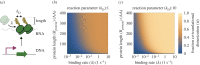
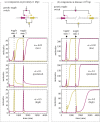
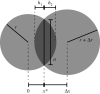
Nonlinearity plays a fundamental role in the performance of both natural and synthetic biological networks. Key functional motifs in living microbial systems, such as the emergence of bistability or oscillations, rely on nonlinear molecular dynamics. Despite its core importance, the rational design of nonlinearity remains an unmet challenge. This is largely due to a lack of mathematical modelling that accounts for the mechanistic basis of nonlinearity. We introduce a model for gene regulatory circuits that explicitly simulates protein dimerization-a well-known source of nonlinear dynamics. Specifically, our approach focuses on modelling co-translational dimerization: the formation of protein dimers during-and not after-translation. This is in contrast to the prevailing assumption that dimer generation is only viable between freely diffusing monomers (i.e. post-translational dimerization). We provide a method for fine-tuning nonlinearity on demand by balancing the impact of co- versus post-translational dimerization. Furthermore, we suggest design rules, such as protein length or physical separation between genes, that may be used to adjust dimerization dynamics in vivo. The design, build and test of genetic circuits with on-demand nonlinear dynamics will greatly improve the programmability of synthetic biological systems.
Keywords: genetic circuits; mathematical modelling; nonlinearity; protein dimerization; synthetic biology; systems biology.
Conflict of interest statement
We declare we have no competing interests.
Figures

Similar articles
-
A Model for the Spatiotemporal Design of Gene Regulatory Circuits †.ACS Synth Biol. 2019 Sep 20;8(9):2007-2016. doi: 10.1021/acssynbio.9b00022. Epub 2019 Aug 30. ACS Synth Biol. 2019. PMID: 31429541
-
A Digitally Programmable Cytomorphic Chip for Simulation of Arbitrary Biochemical Reaction Networks.IEEE Trans Biomed Circuits Syst. 2018 Apr;12(2):360-378. doi: 10.1109/TBCAS.2017.2781253. IEEE Trans Biomed Circuits Syst. 2018. PMID: 29570063 Free PMC article.
-
Complex signal processing in synthetic gene circuits using cooperative regulatory assemblies.Science. 2019 May 10;364(6440):593-597. doi: 10.1126/science.aau8287. Epub 2019 Apr 18. Science. 2019. PMID: 31000590 Free PMC article.
-
Mathematical Modelling in Plant Synthetic Biology.Methods Mol Biol. 2022;2379:209-251. doi: 10.1007/978-1-0716-1791-5_13. Methods Mol Biol. 2022. PMID: 35188665 Review.
-
Synthetic nonlinear computation for genetic circuit design.Curr Opin Biotechnol. 2022 Aug;76:102727. doi: 10.1016/j.copbio.2022.102727. Epub 2022 May 4. Curr Opin Biotechnol. 2022. PMID: 35525177 Review.
Cited by
-
The nonlinearity of regulation in biological networks.NPJ Syst Biol Appl. 2023 Apr 4;9(1):10. doi: 10.1038/s41540-023-00273-w. NPJ Syst Biol Appl. 2023. PMID: 37015937 Free PMC article.
-
An electrogenetic toggle switch model.Microb Biotechnol. 2023 Mar;16(3):546-559. doi: 10.1111/1751-7915.14153. Epub 2022 Oct 7. Microb Biotechnol. 2023. PMID: 36207818 Free PMC article.
-
Deep neural networks with knockoff features identify nonlinear causal relations and estimate effect sizes in complex biological systems.Gigascience. 2022 Dec 28;12:giad044. doi: 10.1093/gigascience/giad044. Epub 2023 Jul 3. Gigascience. 2022. PMID: 37395630 Free PMC article.
References
-
- Amos M, Goñi-Moreno A. 2018. Cellular computing and synthetic biology. In Computational matter (eds S Stepney, S Rasmussen, M Amos), pp. 93–110. New York, NY: Springer.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

