Isobaric Tagging for Relative and Absolute Protein Quantification (iTRAQ)-Based Quantitative Proteomics Analysis of Differentially Expressed Proteins 1 Week After Spinal Cord Injury in a Rat Model
- PMID: 33144554
- PMCID: PMC7650090
- DOI: 10.12659/MSM.924266
Isobaric Tagging for Relative and Absolute Protein Quantification (iTRAQ)-Based Quantitative Proteomics Analysis of Differentially Expressed Proteins 1 Week After Spinal Cord Injury in a Rat Model
Abstract
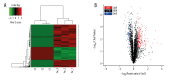
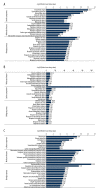
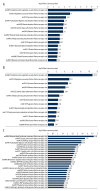
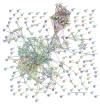
BACKGROUND Spinal cord injury (SCI) is a devastating trauma of the central nervous system (CNS), with high levels of morbidity, disability, and mortality. One week after SCI may be a critical time for treatment. Changes in protein expression have crucial functions in nervous system diseases, although the effects of changes occurring 1 week after SCI on patient outcomes are unclear. MATERIAL AND METHODS Protein expression was examined in a rat contusive SCI model 1 week after SCI. Differentially expressed proteins (DEPs) were identified by isobaric tagging for relative and absolute protein quantification (iTRAQ)-coupled liquid chromatography tandem-mass spectrometry (LC-MS/MS) proteomics analysis. Gene Ontology (GO) analysis was performed to identify the biological processes, molecular functions, and cellular component terms of the identified DEPs, and the Kyoto Encyclopedia of Genes and Genomes (KEGG) was used to identify key enriched pathways. Protein-protein interaction (PPI) networks were analyzed to identify the top 10 high-degree core proteins. RESULTS Of the 295 DEPs identified, 204 (69.15%) were upregulated and 91 (30.85%) were downregulated 1 week after injury. The main cellular components, molecular functions, biological processes, and pathways identified may be crucial mechanisms involved in SCI. The top 10 high-degree core proteins were complement component C3 (C3), alpha-2-HS-glycoprotein (Ahsg), T-kininogen 1 (Kng1), Serpinc1 protein (Serpinc1), apolipoprotein A-I (Apoa1), serum albumin (Alb), disulfide-isomerase protein (P4hb), transport protein Sec61 subunit alpha isoform 1 (Sec61a1), serotransferrin (Tf), and 60S ribosomal protein L15 (Rpl15). CONCLUSIONS The proteins identified in this study may provide potential targets for diagnosis and treatment 1 week after SCI.
Conflict of interest statement
None.
Figures
Similar articles
-
Identification of differentially expressed proteins in rats with spinal cord injury during the transitional phase using an iTRAQ-based quantitative analysis.Gene. 2018 Nov 30;677:66-76. doi: 10.1016/j.gene.2018.07.050. Epub 2018 Jul 20. Gene. 2018. PMID: 30036659
-
Identification of Differentially Expressed Proteins in Rats with Early Subacute Spinal Cord Injury using an iTRAQ-based Quantitative Analysis.Comb Chem High Throughput Screen. 2023;26(11):1960-1973. doi: 10.2174/1386207326666230113152622. Comb Chem High Throughput Screen. 2023. PMID: 36642874
-
A proteomic and phosphoproteomic landscape of spinal cord injury.Neurosci Lett. 2023 Sep 25;814:137449. doi: 10.1016/j.neulet.2023.137449. Epub 2023 Aug 18. Neurosci Lett. 2023. PMID: 37597742
-
Systematic review and meta-analysis of mass spectrometry proteomics applied to ocular fluids to assess potential biomarkers of age-related macular degeneration.BMC Ophthalmol. 2023 Dec 12;23(1):507. doi: 10.1186/s12886-023-03237-0. BMC Ophthalmol. 2023. PMID: 38087257 Free PMC article.
-
Isobaric protein and peptide quantification: perspectives and issues.Expert Rev Proteomics. 2010 Oct;7(5):647-53. doi: 10.1586/epr.10.29. Expert Rev Proteomics. 2010. PMID: 20973638 Review.
Cited by
-
The role of foam cells in spinal cord injury: challenges and opportunities for intervention.Front Immunol. 2024 Mar 13;15:1368203. doi: 10.3389/fimmu.2024.1368203. eCollection 2024. Front Immunol. 2024. PMID: 38545108 Free PMC article. Review.
-
Complement Receptor 3 Pathway and NMDA Receptor 2B Subunit Involve Neuropathic Pain Associated with Spinal Cord Injury.J Pain Res. 2022 Jun 25;15:1813-1823. doi: 10.2147/JPR.S366782. eCollection 2022. J Pain Res. 2022. PMID: 35784110 Free PMC article.
References
-
- Kang Y, Ding H, Zhou HX, et al. Epidemiology of worldwide spinal cord injury: A literature review. J Neurorestoratology. 2018;6:1–9.
-
- Ning GZ, Yu TQ, Feng SQ, et al. Epidemiology of traumatic spinal cord injury in Tianjin, China. Spinal Cord. 2011;49(3):386–90. - PubMed
-
- Ahuja CS, Wilson JR, Nori S, et al. Traumatic spinal cord injury. Nat Rev Dis Primers. 2017;3:17018. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

