Mito Hacker: a set of tools to enable high-throughput analysis of mitochondrial network morphology
- PMID: 33144635
- PMCID: PMC7642274
- DOI: 10.1038/s41598-020-75899-5
Mito Hacker: a set of tools to enable high-throughput analysis of mitochondrial network morphology
Abstract
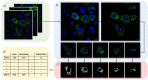
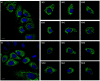
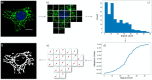
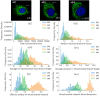
Mitochondria are highly dynamic organelles that can exhibit a wide range of morphologies. Mitochondrial morphology can differ significantly across cell types, reflecting different physiological needs, but can also change rapidly in response to stress or the activation of signaling pathways. Understanding both the cause and consequences of these morphological changes is critical to fully understanding how mitochondrial function contributes to both normal and pathological physiology. However, while robust and quantitative analysis of mitochondrial morphology has become increasingly accessible, there is a need for new tools to generate and analyze large data sets of mitochondrial images in high throughput. The generation of such datasets is critical to fully benefit from rapidly evolving methods in data science, such as neural networks, that have shown tremendous value in extracting novel biological insights and generating new hypotheses. Here we describe a set of three computational tools, Cell Catcher, Mito Catcher and MiA, that we have developed to extract extensive mitochondrial network data on a single-cell level from multi-cell fluorescence images. Cell Catcher automatically separates and isolates individual cells from multi-cell images; Mito Catcher uses the statistical distribution of pixel intensities across the mitochondrial network to detect and remove background noise from the cell and segment the mitochondrial network; MiA uses the binarized mitochondrial network to perform more than 100 mitochondria-level and cell-level morphometric measurements. To validate the utility of this set of tools, we generated a database of morphological features for 630 individual cells that encode 0, 1 or 2 alleles of the mitochondrial fission GTPase Drp1 and demonstrate that these mitochondrial data could be used to predict Drp1 genotype with 87% accuracy. Together, this suite of tools enables the high-throughput and automated collection of detailed and quantitative mitochondrial structural information at a single-cell level. Furthermore, the data generated with these tools, when combined with advanced data science approaches, can be used to generate novel biological insights.
Conflict of interest statement
The authors declare no competing interests.
Figures


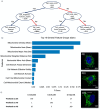
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

