Pathway enrichment analysis of virus-host interactome and prioritization of novel compounds targeting the spike glycoprotein receptor binding domain-human angiotensin-converting enzyme 2 interface to combat SARS-CoV-2
- PMID: 33146070
- PMCID: PMC7651197
- DOI: 10.1080/07391102.2020.1841681
Pathway enrichment analysis of virus-host interactome and prioritization of novel compounds targeting the spike glycoprotein receptor binding domain-human angiotensin-converting enzyme 2 interface to combat SARS-CoV-2
Abstract
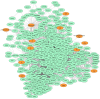
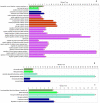
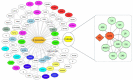
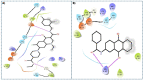
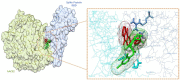
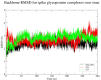
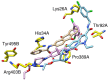
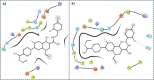
SARS-CoV-2 has become a pandemic causing a serious global health concern. The absence of effective drugs for treatment of the disease has caused its rapid spread on a global scale. Similarly to the SARS-CoV, the SARS-CoV-2 is also involved in a complex interplay with the host cells. This infection is characterized by a diffused alveolar damage consistent with the Acute Respiratory Disease Syndrome (ARDS). To explore the complex mechanisms of the disease at the system level, we used a network medicine tools approach. The protein-protein interactions (PPIs) between the SARS-CoV and the associated human cell proteins are crucial for the viral pathogenesis. Since the cellular entry of SARS-CoV-2 is accomplished by binding of the spike glycoprotein binding domain (RBD) to the human angiotensin-converting enzyme 2 (hACE2), a molecule that can bind to the spike RDB-hACE2 interface could block the virus entry. Here, we performed a virtual screening of 55 compounds to identify potential molecules that can bind to the spike glycoprotein and spike-ACE2 complex interface. It was found that the compound ethyl 1-{3-[(2,4-dichlorobenzyl) carbamoyl]-1-ethyl-6-fluoro-4-oxo-1,4-dihydro-7-quinolinyl}-4-piperidine carboxylate (the S54 ligand) and ethyl 1-{3-[(2,4-dichlorobenzyl) carbamoyl]-1-ethyl-6-fluoro-4-oxo-1,4-dihydro-7-quinolinyl}-4 piperazine carboxylate (the S55 ligand) forms hydrophobic interactions with Tyr41A, Tyr505B and Tyr553B, Leu29A, Phe495B, respectively of the spike glycoprotein, the hotspot residues in the spike glycoprotein RBD-hACE2 binding interface. Furthermore, molecular dynamics simulations and free energy calculations using the MM-GBSA method showed that the S54 ligand is a stronger binder than a known SARS-CoV spike inhibitor SSAA09E3 (N-(9,10-dioxo-9, 10-dihydroanthracen-2-yl) benzamide).Communicated by Ramaswamy H. Sarma.
Keywords: SARS-CoV-2; molecular docking; molecular dynamics; protein–protein interactions (PPIs); spike glycoprotein; virtual screening.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures


Similar articles
-
Inhibition of S-protein RBD and hACE2 Interaction for Control of SARSCoV- 2 Infection (COVID-19).Mini Rev Med Chem. 2021;21(6):689-703. doi: 10.2174/1389557520666201117111259. Mini Rev Med Chem. 2021. PMID: 33208074 Review.
-
Identification of potential SARS-CoV-2 entry inhibitors by targeting the interface region between the spike RBD and human ACE2.J Infect Public Health. 2021 Feb;14(2):227-237. doi: 10.1016/j.jiph.2020.12.014. Epub 2020 Dec 21. J Infect Public Health. 2021. PMID: 33493919 Free PMC article.
-
De novo design, retrosynthetic analysis and combinatorial synthesis of a hybrid antiviral (VTAR-01) to inhibit the interaction of SARS-CoV2 spike glycoprotein with human angiotensin-converting enzyme 2.Biol Open. 2020 Oct 15;9(10):bio054056. doi: 10.1242/bio.054056. Biol Open. 2020. PMID: 32878881 Free PMC article.
-
Multidisciplinary Approaches Identify Compounds that Bind to Human ACE2 or SARS-CoV-2 Spike Protein as Candidates to Block SARS-CoV-2-ACE2 Receptor Interactions.mBio. 2021 Mar 30;12(2):e03681-20. doi: 10.1128/mBio.03681-20. mBio. 2021. PMID: 33785634 Free PMC article.
-
The expression of hACE2 receptor protein and its involvement in SARS-CoV-2 entry, pathogenesis, and its application as potential therapeutic target.Tumour Biol. 2021;43(1):177-196. doi: 10.3233/TUB-200084. Tumour Biol. 2021. PMID: 34420993 Review.
Cited by
-
Nucleocapsid and Spike Proteins of SARS-CoV-2 Drive Neutrophil Extracellular Trap Formation.Immune Netw. 2021 Feb 23;21(2):e16. doi: 10.4110/in.2021.21.e16. eCollection 2021 Apr. Immune Netw. 2021. PMID: 33996172 Free PMC article.
-
In silico binding profile characterization of SARS-CoV-2 spike protein and its mutants bound to human ACE2 receptor.Brief Bioinform. 2021 Nov 5;22(6):bbab188. doi: 10.1093/bib/bbab188. Brief Bioinform. 2021. PMID: 34013346 Free PMC article.
-
A computational analysis of molecular evolution for virulence genes of zoonotic novel coronavirus (COVID-19).Comput Biol Chem. 2021 Aug;93:107532. doi: 10.1016/j.compbiolchem.2021.107532. Epub 2021 Jun 19. Comput Biol Chem. 2021. PMID: 34171504 Free PMC article.
-
The effect of quercetin supplementation on clinical outcomes in COVID-19 patients: A systematic review and meta-analysis.Food Sci Nutr. 2023 Sep 26;11(12):7504-7514. doi: 10.1002/fsn3.3715. eCollection 2023 Dec. Food Sci Nutr. 2023. PMID: 38107099 Free PMC article. Review.
-
Establishment of a Yeast Two-Hybrid-Based High-Throughput Screening Model for Selection of SARS-CoV-2 Spike-ACE2 Interaction Inhibitors.Int J Mol Sci. 2025 Jan 15;26(2):678. doi: 10.3390/ijms26020678. Int J Mol Sci. 2025. PMID: 39859397 Free PMC article.
References
-
- Abdelli, I., Hassani, F., Bekkel Brikci, S., & Ghalem, S. (2020). In silico study the inhibition of angiotensin converting enzyme 2 receptor of COVID-19 by Ammoides verticillata components harvested from Western Algeria. Journal of Biomolecular Structure and Dynamics, 1–14. 10.1080/07391102.2020.1763199 - DOI - PMC - PubMed
-
- Albert, R., & Barabási, A. L. (2002). Statistical mechanics of complex networks. Reviews of Modern Physics, 74(1), 47–97. 10.1103/RevModPhys.74.47. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
