Temporal Quantitative Profiling of Newly Synthesized Proteins during Aβ Accumulation
- PMID: 33147027
- PMCID: PMC7775911
- DOI: 10.1021/acs.jproteome.0c00645
Temporal Quantitative Profiling of Newly Synthesized Proteins during Aβ Accumulation
Abstract
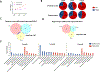
Accumulation of aggregated amyloid beta (Aβ) in the brain is believed to impair multiple cellular pathways and play a central role in Alzheimer's disease pathology. However, how this process is regulated remains unclear. In theory, measuring protein synthesis is the most direct way to evaluate a cell's response to stimuli, but to date, there have been few reliable methods to do this. To identify the protein regulatory network during the development of Aβ deposition in AD, we applied a new proteomic technique to quantitate newly synthesized protein (NSP) changes in the cerebral cortex and hippocampus of 2-, 5-, and 9-month-old APP/PS1 AD transgenic mice. This bio-orthogonal noncanonical amino acid tagging analysis combined PALM (pulse azidohomoalanine labeling in mammals) and HILAQ (heavy isotope labeled AHA quantitation) to reveal a comprehensive dataset of NSPs prior to and post Aβ deposition, including the identification of proteins not previously associated with AD, and demonstrated that the pattern of differentially expressed NSPs is age-dependent. We also found dysregulated vesicle transportation networks including endosomal subunits, coat protein complex I (COPI), and mitochondrial respiratory chain throughout all time points and two brain regions. These results point to a pathological dysregulation of vesicle transportation which occurs prior to Aβ accumulation and the onset of AD symptoms, which may progressively impact the entire protein network and thereby drive neurodegeneration. This study illustrates key pathway regulation responses to the development of AD pathogenesis by directly measuring the changes in protein synthesis and provides unique insights into the mechanisms that underlie AD.
Keywords: Alzheimer’s disease; Aβ accumulation; newly synthesized proteins; quantitative profiling.
Figures

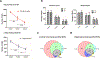




Similar articles
-
Abnormal interaction of VDAC1 with amyloid beta and phosphorylated tau causes mitochondrial dysfunction in Alzheimer's disease.Hum Mol Genet. 2012 Dec 1;21(23):5131-46. doi: 10.1093/hmg/dds360. Epub 2012 Aug 27. Hum Mol Genet. 2012. PMID: 22926141 Free PMC article.
-
Early Presymptomatic Changes in the Proteome of Mitochondria-Associated Membrane in the APP/PS1 Mouse Model of Alzheimer's Disease.Mol Neurobiol. 2018 Oct;55(10):7839-7857. doi: 10.1007/s12035-018-0955-6. Epub 2018 Feb 22. Mol Neurobiol. 2018. PMID: 29468564
-
β-Amyloid, cholinergic transmission, and cerebrovascular system -- a developmental study in a mouse model of Alzheimer's disease.Curr Pharm Des. 2013;19(38):6749-65. doi: 10.2174/13816128113199990711. Curr Pharm Des. 2013. PMID: 23530514
-
Alzheimer's disease.Subcell Biochem. 2012;65:329-52. doi: 10.1007/978-94-007-5416-4_14. Subcell Biochem. 2012. PMID: 23225010 Review.
-
APP transgenic modeling of Alzheimer's disease: mechanisms of neurodegeneration and aberrant neurogenesis.Brain Struct Funct. 2010 Mar;214(2-3):111-26. doi: 10.1007/s00429-009-0232-6. Epub 2009 Nov 29. Brain Struct Funct. 2010. PMID: 20091183 Free PMC article. Review.
Cited by
-
Age-dependent shift in the de novo proteome accompanies pathogenesis in an Alzheimer's disease mouse model.Commun Biol. 2021 Jun 30;4(1):823. doi: 10.1038/s42003-021-02324-6. Commun Biol. 2021. PMID: 34193971 Free PMC article.
-
Light-Activatable, Cell-Type Specific Labeling of the Nascent Proteome.ACS Chem Neurosci. 2024 Oct 2;15(19):3473-3481. doi: 10.1021/acschemneuro.4c00274. Epub 2024 Sep 22. ACS Chem Neurosci. 2024. PMID: 39307974 Free PMC article.
-
Mutations in the COPI coatomer subunit α-COP induce release of Aβ-42 and amyloid precursor protein intracellular domain and increase tau oligomerization and release.Neurobiol Aging. 2021 May;101:57-69. doi: 10.1016/j.neurobiolaging.2021.01.003. Epub 2021 Jan 13. Neurobiol Aging. 2021. PMID: 33582567 Free PMC article.
-
Proteomics of the heart.Physiol Rev. 2024 Jul 1;104(3):931-982. doi: 10.1152/physrev.00026.2023. Epub 2024 Feb 1. Physiol Rev. 2024. PMID: 38300522 Free PMC article. Review.
-
In-Depth Proteomic Analysis of De Novo Proteome in a Mouse Model of Alzheimer's Disease.J Alzheimers Dis. 2023;91(4):1471-1482. doi: 10.3233/JAD-221044. J Alzheimers Dis. 2023. PMID: 36641677 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

