LINC00968 can inhibit the progression of lung adenocarcinoma through the miR-21-5p/SMAD7 signal axis
- PMID: 33147570
- PMCID: PMC7695398
- DOI: 10.18632/aging.104011
LINC00968 can inhibit the progression of lung adenocarcinoma through the miR-21-5p/SMAD7 signal axis
Abstract
Background: Long non-coding RNAs (LncRNAs) have been associated with several types of cancer. However, little is known about their role in lung adenocarcinoma (LUAD).
Results: LINC00968 was significantly differentially expressed in LUAD tissues. Downregulated LINC00968 was associated with clinicopathological features of LUAD. LINC00968 inhibited cell growth and metastasis by regulating the Hippo signaling pathway We demonstrated that LINC00968 acts as a ceRNA to consume miR-21-5p, enhancing the accumulation of SMAD7, a miR-21-5p target.
Conclusions: LINC00968 limits LUAD progression via the miR-21-5p/SMAD7 axis and may serve as a prognostic biomarker and therapeutic target for LUAD.
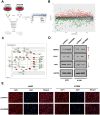
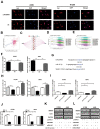
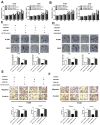
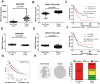
Methods: We conducted comprehensive data mining on LINC00968 based on the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) database. The expression of LINC00968 in LUAD cells was determined using in situ hybridization. We detected LINC00968 function in LUAD cells using the MTT, clone formation, and transwell assays, and tumor xenografts. Label-free quantitative proteomics, western blotting, a dual-luciferase reporter assay, immunofluorescence, and RNA immunoprecipitation assays were used to determine the correlations among LINC00968, miR-21-5p, and SMAD7. Gain- and loss-function approaches were used to explore the effects of LINC00968, miR-21-5p, and SMAD7 on cell proliferation, migration, and invasion.
Keywords: LINC00968; SMAD7; lung adenocarcinoma (LUAD); metastasis; prognosis.
Conflict of interest statement
Figures


Similar articles
-
linc00968 inhibits the tumorigenesis and metastasis of lung adenocarcinoma via serving as a ceRNA against miR-9-5p and increasing CPEB3.Aging (Albany NY). 2020 Nov 5;12(22):22582-22598. doi: 10.18632/aging.103833. Epub 2020 Nov 5. Aging (Albany NY). 2020. PMID: 33159015 Free PMC article.
-
HCG18/miR-34a-5p/HMMR axis accelerates the progression of lung adenocarcinoma.Biomed Pharmacother. 2020 Sep;129:110217. doi: 10.1016/j.biopha.2020.110217. Epub 2020 Jun 16. Biomed Pharmacother. 2020. PMID: 32559619
-
USF1-induced overexpression of long noncoding RNA WDFY3-AS2 promotes lung adenocarcinoma progression via targeting miR-491-5p/ZNF703 axis.Mol Carcinog. 2020 Aug;59(8):875-885. doi: 10.1002/mc.23181. Epub 2020 Apr 10. Mol Carcinog. 2020. PMID: 32275336
-
MicroRNA‑mediated regulation in lung adenocarcinoma: Signaling pathways and potential therapeutic implications (Review).Oncol Rep. 2023 Dec;50(6):211. doi: 10.3892/or.2023.8648. Epub 2023 Oct 20. Oncol Rep. 2023. PMID: 37859595 Free PMC article. Review.
-
miR-181a/b therapy in lung cancer: reality or myth?Mol Oncol. 2019 Jan;13(1):9-25. doi: 10.1002/1878-0261.12420. Epub 2019 Jan 3. Mol Oncol. 2019. PMID: 30548184 Free PMC article. Review.
Cited by
-
Identification of a Ferroptosis-Related LncRNA Signature as a Novel Prognosis Model for Lung Adenocarcinoma.Front Oncol. 2021 Jun 23;11:675545. doi: 10.3389/fonc.2021.675545. eCollection 2021. Front Oncol. 2021. PMID: 34249715 Free PMC article.
-
microRNA 21 and long non-coding RNAs interplays underlie cancer pathophysiology: A narrative review.Noncoding RNA Res. 2024 Mar 31;9(3):831-852. doi: 10.1016/j.ncrna.2024.03.013. eCollection 2024 Sep. Noncoding RNA Res. 2024. PMID: 38586315 Free PMC article. Review.
-
Hypoxic Tumor-Derived Exosomal miR-199a-3p Promote Gastric Cancer Metastasis via MAP3K4.J Cancer. 2023 Jul 15;14(11):2161-2172. doi: 10.7150/jca.83909. eCollection 2023. J Cancer. 2023. PMID: 37497404 Free PMC article.
-
Long noncoding RNA LINC00968 inhibits proliferation, migration and invasion of lung adenocarcinoma through targeting miR-22-5p/CDC14A axis.3 Biotech. 2021 Oct;11(10):433. doi: 10.1007/s13205-021-02981-8. Epub 2021 Sep 14. 3 Biotech. 2021. Retraction in: 3 Biotech. 2023 May;13(5):120. doi: 10.1007/s13205-023-03531-0. PMID: 34603911 Free PMC article. Retracted.
-
Regulation of TGFβ/SMAD signaling by long non-coding RNAs in different cancers: Dark Knight in the Castle of molecular oncology.Noncoding RNA Res. 2021 Jan 7;6(1):23-28. doi: 10.1016/j.ncrna.2020.12.003. eCollection 2021 Mar. Noncoding RNA Res. 2021. PMID: 33511320 Free PMC article.
References
-
- Higuchi T, Oshiro H, Zhang Z, Miyake K, Sugisawa N, Katsuya Y, Yamamoto N, Hayashi K, Kimura H, Miwa S, Igarashi K, Zhao M, Bouvet M, et al.. Osimertinib regresses an EGFR-mutant cisplatinum- resistant lung adenocarcinoma growing in the brain in nude mice. Transl Oncol. 2019; 12:640–45. 10.1016/j.tranon.2019.01.007 - DOI - PMC - PubMed
-
- Agirre X, Meydan C, Jiang Y, Garate L, Doane AS, Li Z, Verma A, Paiva B, Martín-Subero JI, Elemento O, Mason CE, Prosper F, Melnick A. Long non-coding RNAs discriminate the stages and gene regulatory states of human humoral immune response. Nat Commun. 2019; 10:821. 10.1038/s41467-019-08679-z - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

