Functional Analyses of Bovine Foamy Virus-Encoded miRNAs Reveal the Importance of a Defined miRNA for Virus Replication and Host-Virus Interaction
- PMID: 33147813
- PMCID: PMC7693620
- DOI: 10.3390/v12111250
Functional Analyses of Bovine Foamy Virus-Encoded miRNAs Reveal the Importance of a Defined miRNA for Virus Replication and Host-Virus Interaction
Abstract
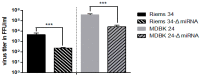
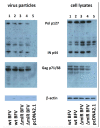
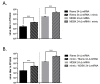
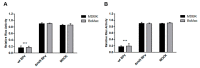
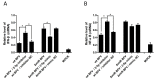
In addition to regulatory or accessory proteins, some complex retroviruses gain a repertoire of micro-RNAs (miRNAs) to regulate and control virus-host interactions for efficient replication and spread. In particular, bovine and simian foamy viruses (BFV and SFV) have recently been shown to express a diverse set of RNA polymerase III-directed miRNAs, some with a unique primary miRNA double-hairpin, dumbbell-shaped structure not known in other viruses or organisms. While the mechanisms of expression and structural requirements have been studied, the functional importance of these miRNAs is still far from understood. Here, we describe the in silico identification of BFV miRNA targets and the subsequent experimental validation of bovine Ankyrin Repeat Domain 17 (ANKRD17) and Bax-interacting factor 1 (Bif1) target genes in vitro and, finally, the suppression of ANKRD17 downstream genes in the affected pathway. Deletion of the entire miRNA cassette in the non-coding part of the U3 region of the long terminal repeats attenuated replication of corresponding BFV mutants in bovine cells. This repression can be almost completely trans-complemented by the most abundant miRNA BF2-5p having the best scores for predicted and validated BFV miRNA target genes. Deletion of the miRNA cassette does not grossly affect particle release and overall particle composition.
Keywords: ANKRD17; Bif1 (SH3GLB1); bovine foamy virus; innate immunity; miRNA expression; miRNA target gene identification; replication in vitro; spumaretrovirus; virus-host-interaction.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

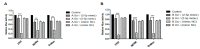
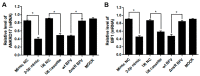
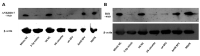

Similar articles
-
Functional characterization of the bovine foamy virus miRNA expression cassette and its dumbbell-shaped pri-miRNA.Virus Genes. 2018 Aug;54(4):550-560. doi: 10.1007/s11262-018-1574-z. Epub 2018 May 31. Virus Genes. 2018. PMID: 29855776
-
Identification of novel, highly expressed retroviral microRNAs in cells infected by bovine foamy virus.J Virol. 2014 May;88(9):4679-86. doi: 10.1128/JVI.03587-13. Epub 2014 Feb 12. J Virol. 2014. PMID: 24522910 Free PMC article.
-
Bovine Foamy Virus: Shared and Unique Molecular Features In Vitro and In Vivo.Viruses. 2019 Nov 21;11(12):1084. doi: 10.3390/v11121084. Viruses. 2019. PMID: 31766538 Free PMC article. Review.
-
Noncanonical microRNA (miRNA) biogenesis gives rise to retroviral mimics of lymphoproliferative and immunosuppressive host miRNAs.mBio. 2014 Apr 8;5(2):e00074. doi: 10.1128/mBio.00074-14. mBio. 2014. PMID: 24713319 Free PMC article.
-
Foamy virus vectors: an awaited alternative to gammaretro- and lentiviral vectors.Curr Gene Ther. 2007 Aug;7(4):261-71. doi: 10.2174/156652307781369092. Curr Gene Ther. 2007. PMID: 17969559 Review.
Cited by
-
Emerging delivery approaches for targeted pulmonary fibrosis treatment.Adv Drug Deliv Rev. 2024 Jan;204:115147. doi: 10.1016/j.addr.2023.115147. Epub 2023 Dec 6. Adv Drug Deliv Rev. 2024. PMID: 38065244 Free PMC article. Review.
-
Molecular Epidemiology and Whole-Genome Analysis of Bovine Foamy Virus in Japan.Viruses. 2021 May 28;13(6):1017. doi: 10.3390/v13061017. Viruses. 2021. PMID: 34071542 Free PMC article.
-
RNA-based therapies: A cog in the wheel of lung cancer defense.Mol Cancer. 2021 Mar 19;20(1):54. doi: 10.1186/s12943-021-01338-2. Mol Cancer. 2021. PMID: 33740988 Free PMC article. Review.
-
The Unique, the Known, and the Unknown of Spumaretrovirus Assembly.Viruses. 2021 Jan 13;13(1):105. doi: 10.3390/v13010105. Viruses. 2021. PMID: 33451128 Free PMC article. Review.
-
An Outbred Calf Model for Determining Innate Immune Sensing and Evolutionary Trajectories of a Cell Culture-Adapted Bovine Foamy Virus Variant.Viruses. 2023 Aug 20;15(8):1772. doi: 10.3390/v15081772. Viruses. 2023. PMID: 37632114 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

