Genome-scale phylogeny and contrasting modes of genome evolution in the fungal phylum Ascomycota
- PMID: 33148650
- PMCID: PMC7673691
- DOI: 10.1126/sciadv.abd0079
Genome-scale phylogeny and contrasting modes of genome evolution in the fungal phylum Ascomycota
Abstract
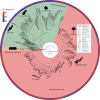
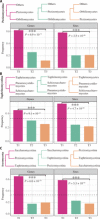
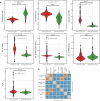
Ascomycota, the largest and most well-studied phylum of fungi, contains three subphyla: Saccharomycotina (budding yeasts), Pezizomycotina (filamentous fungi), and Taphrinomycotina (fission yeasts). Despite its importance, we lack a comprehensive genome-scale phylogeny or understanding of the similarities and differences in the mode of genome evolution within this phylum. By examining 1107 genomes from Saccharomycotina (332), Pezizomycotina (761), and Taphrinomycotina (14) species, we inferred a robust genome-wide phylogeny that resolves several contentious relationships and estimated that the Ascomycota last common ancestor likely originated in the Ediacaran period. Comparisons of genomic properties revealed that Saccharomycotina and Pezizomycotina differ greatly in their genome properties and enabled inference of the direction of evolutionary change. The Saccharomycotina typically have smaller genomes, lower guanine-cytosine contents, lower numbers of genes, and higher rates of molecular sequence evolution compared with Pezizomycotina. These results provide a robust evolutionary framework for understanding the diversity and ecological lifestyles of the largest fungal phylum.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works. Distributed under a Creative Commons Attribution NonCommercial License 4.0 (CC BY-NC).
Figures

Similar articles
-
Comparison of protein coding gene contents of the fungal phyla Pezizomycotina and Saccharomycotina.BMC Genomics. 2007 Sep 17;8:325. doi: 10.1186/1471-2164-8-325. BMC Genomics. 2007. PMID: 17868481 Free PMC article.
-
Evolution of SET-domain protein families in the unicellular and multicellular Ascomycota fungi.BMC Evol Biol. 2008 Jul 1;8:190. doi: 10.1186/1471-2148-8-190. BMC Evol Biol. 2008. PMID: 18593478 Free PMC article.
-
Phylogenomic analyses support the monophyly of Taphrinomycotina, including Schizosaccharomyces fission yeasts.Mol Biol Evol. 2009 Jan;26(1):27-34. doi: 10.1093/molbev/msn221. Epub 2008 Oct 14. Mol Biol Evol. 2009. PMID: 18922765 Free PMC article.
-
Evolution of Mating in the Saccharomycotina.Annu Rev Microbiol. 2017 Sep 8;71:197-214. doi: 10.1146/annurev-micro-090816-093403. Epub 2017 Jun 28. Annu Rev Microbiol. 2017. PMID: 28657889 Review.
-
The Fungal Tree of Life: from Molecular Systematics to Genome-Scale Phylogenies.Microbiol Spectr. 2017 Sep;5(5):10.1128/microbiolspec.funk-0053-2016. doi: 10.1128/microbiolspec.FUNK-0053-2016. Microbiol Spectr. 2017. PMID: 28917057 Free PMC article. Review.
Cited by
-
Giant Starship Elements Mobilize Accessory Genes in Fungal Genomes.Mol Biol Evol. 2022 May 3;39(5):msac109. doi: 10.1093/molbev/msac109. Mol Biol Evol. 2022. PMID: 35588244 Free PMC article.
-
Coevolutionary insights between promoters and transcription factors in the plant and animal kingdoms.Zool Res. 2022 Sep 18;43(5):805-812. doi: 10.24272/j.issn.2095-8137.2022.111. Zool Res. 2022. PMID: 35993132 Free PMC article.
-
Functional Clustering of Metabolically Related Genes Is Conserved across Dikarya.J Fungi (Basel). 2023 Apr 28;9(5):523. doi: 10.3390/jof9050523. J Fungi (Basel). 2023. PMID: 37233234 Free PMC article. Review.
-
Whole-genome based phylogeny and comparative genomics of Sporidiobolales and related taxa of Basidiomycetes.IMA Fungus. 2025 May 13;16:e141626. doi: 10.3897/imafungus.16.141626. eCollection 2025. IMA Fungus. 2025. PMID: 40400710 Free PMC article.
-
Membrane and organelle rearrangement during ascospore formation in budding yeast.Microbiol Mol Biol Rev. 2024 Sep 26;88(3):e0001324. doi: 10.1128/mmbr.00013-24. Epub 2024 Jun 20. Microbiol Mol Biol Rev. 2024. PMID: 38899894 Free PMC article. Review.
References
-
- James T. Y., Stajich J. E., Hittinger C. T., Rokas A., Toward a fully resolved fungal tree of life. Annu. Rev. Microbiol. 74, 291–313 (2020). - PubMed
-
- Shen X.-X., Opulente D. A., Kominek J., Zhou X., Steenwyk J. L., Buh K. V., Haase M. A. B., Wisecaver J. H., Wang M., Doering D. T., Boudouris J. T., Schneider R. M., Langdon Q. K., Ohkuma M., Endoh R., Takashima M., Manabe R.-i., Čadež N., Libkind D., Rosa C. A., DeVirgilio J., Hulfachor A. B., Groenewald M., Kurtzman C. P., Hittinger C. T., Rokas A., Tempo and mode of genome evolution in the budding yeast subphylum. Cell 175, 1533–1545.e20 (2018). - PMC - PubMed
-
- Peter J., De Chiara M., Friedrich A., Yue J.-X., Pflieger D., Bergström A., Sigwalt A., Barre B., Freel K., Llored A., Cruaud C., Labadie K., Aury J.-M., Istace B., Lebrigand K., Barbry P., Engelen S., Lemainque A., Wincker P., Liti G., Schacherer J., Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556, 339–344 (2018). - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

