Host variables confound gut microbiota studies of human disease
- PMID: 33149306
- PMCID: PMC7677204
- DOI: 10.1038/s41586-020-2881-9
Host variables confound gut microbiota studies of human disease
Abstract
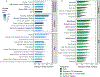
Low concordance between studies that examine the role of microbiota in human diseases is a pervasive challenge that limits the capacity to identify causal relationships between host-associated microorganisms and pathology. The risk of obtaining false positives is exacerbated by wide interindividual heterogeneity in microbiota composition1, probably due to population-wide differences in human lifestyle and physiological variables2 that exert differential effects on the microbiota. Here we infer the greatest, generalized sources of heterogeneity in human gut microbiota profiles and also identify human lifestyle and physiological characteristics that, if not evenly matched between cases and controls, confound microbiota analyses to produce spurious microbial associations with human diseases. We identify alcohol consumption frequency and bowel movement quality as unexpectedly strong sources of gut microbiota variance that differ in distribution between healthy participants and participants with a disease and that can confound study designs. We demonstrate that for numerous prevalent, high-burden human diseases, matching cases and controls for confounding variables reduces observed differences in the microbiota and the incidence of spurious associations. On this basis, we present a list of host variables that we recommend should be captured in human microbiota studies for the purpose of matching comparison groups, which we anticipate will increase robustness and reproducibility in resolving the members of the gut microbiota that are truly associated with human disease.
Conflict of interest statement
Competing Interests
R.K. is a director of the Center for Microbiome Innovation at UC San Diego, which receives industry research funding for various microbiome initiatives, but no industry funding was provided for this project. The remaining authors declare no competing interests.
Figures
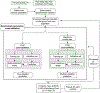

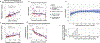


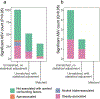

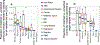

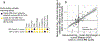
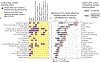
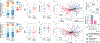

Comment in
-
Identifying gut microbes that affect human health.Nature. 2020 Nov;587(7834):373-374. doi: 10.1038/d41586-020-03069-8. Nature. 2020. PMID: 33149313 No abstract available.
-
Avoiding the pitfalls in microbiota studies.Nat Rev Microbiol. 2021 Jan;19(1):2. doi: 10.1038/s41579-020-00480-w. Nat Rev Microbiol. 2021. PMID: 33154571 No abstract available.

