Genome-wide analysis of acute traumatic spinal cord injury-related RNA expression profiles and uncovering of a regulatory axis in spinal fibrotic scars
- PMID: 33150698
- PMCID: PMC7791181
- DOI: 10.1111/cpr.12951
Genome-wide analysis of acute traumatic spinal cord injury-related RNA expression profiles and uncovering of a regulatory axis in spinal fibrotic scars
Abstract
Objectives: Long non-coding RNAs (lncRNAs) are critical for posttranscriptional and transcriptional regulation in eukaryotic cells. However, data on lncRNA expression in the lesion epicentres of spinal tissues after acute traumatic spinal cord injury (ATSCI) are scarce. We aimed to identify lncRNA expression profiles in such centres and predict latent regulatory networks.
Materials and methods: High-throughput RNA-sequencing was used to profile the expression and regulatory patterns of lncRNAs, microRNAs and messenger RNAs (mRNAs) in an ATSCI C57BL/6 mouse model. Chromosome distributions, open reading frames (ORFs), transcript abundances, exon numbers and lengths were compared between lncRNAs and mRNAs. Gene ontology, KEGG pathways and binding networks were analysed. The findings were validated by qRT-PCRs and luciferase assays.
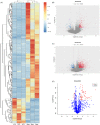
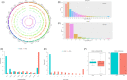
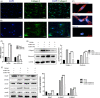
Results: Intronic lncRNAs were the most common differentially expressed lncRNA. Most lncRNAs had <6 exons, and lncRNAs had shorter lengths and lesser ORFs than mRNAs. MiR-21a-5p had the most significant differential expression and bound to the differentially expressed lncRNA ENSMUST00000195880. The microRNAs and lncRNAs with significant differential expression were screened, and a lncRNA/miRNA/mRNA interaction network was predicted, constructed and verified.
Conclusions: The regulatory actions of this network may play a role in the pathophysiology of ATSCI. Our findings may lead to better understanding of potential ncRNA biomarkers and confer better therapeutic strategies for ATSCIs.
Keywords: acute traumatic spinal cord injury; interaction network; long non-coding RNAs; mRNA; miRNA.
© 2020 The Authors. Cell Proliferation Published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interests. The sponsors had no role in the design, execution, interpretation or writing of the study, or in the decision to publish the results.
Figures



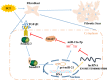
References
-
- Harvey LA, Dunlop SA, Churilov L, Galea MP. Early intensive hand rehabilitation is not more effective than usual care plus one‐to‐one hand therapy in people with sub‐acute spinal cord injury ('Hands On'): a randomised trial. J Physiother. 2017;63(4):197‐204. - PubMed
-
- Hara M, Kobayakawa K, Ohkawa Y, et al. Interaction of reactive astrocytes with type I collagen induces astrocytic scar formation through the integrin‐N‐cadherin pathway after spinal cord injury. Nat Med. 2017;23(7):818‐828. - PubMed
MeSH terms
Substances
Grants and funding
- NO.2019JDRC0100/The Innovation and Entrepreneurship Project of Sichuan Technology Gallery
- NO.20MZGC0265/The Innovation and Entrepreneurship Project of Sichuan Technology Gallery
- Z20192013/the National Clinical Research Center for Geriatrics, West China Hospital, Sichuan University
- Y2018B22/the National Clinical Research Center for Geriatrics, West China Hospital, Sichuan University
- NO.2018SZ0159/the Science and Technology Support Project of Sichuan Province
LinkOut - more resources
Full Text Sources
Medical

