Transporters Involved in the Biogenesis and Functionalization of the Mycobacterial Cell Envelope
- PMID: 33170669
- PMCID: PMC8107195
- DOI: 10.1021/acs.chemrev.0c00869
Transporters Involved in the Biogenesis and Functionalization of the Mycobacterial Cell Envelope
Abstract
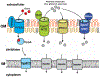
The biology of mycobacteria is dominated by a complex cell envelope of unique composition and structure and of exceptionally low permeability. This cell envelope is the basis of many of the pathogenic features of mycobacteria and the site of susceptibility and resistance to many antibiotics and host defense mechanisms. This review is focused on the transporters that assemble and functionalize this complex structure. It highlights both the progress and the limits of our understanding of how (lipo)polysaccharides, (glyco)lipids, and other bacterial secretion products are translocated across the different layers of the cell envelope to their final extra-cytoplasmic location. It further describes some of the unique strategies evolved by mycobacteria to import nutrients and other products through this highly impermeable barrier.
Figures







Similar articles
-
Transport of outer membrane lipids in mycobacteria.Biochim Biophys Acta Mol Cell Biol Lipids. 2017 Nov;1862(11):1340-1354. doi: 10.1016/j.bbalip.2017.01.005. Epub 2017 Jan 19. Biochim Biophys Acta Mol Cell Biol Lipids. 2017. PMID: 28110100 Review.
-
The envelope of mycobacteria.Annu Rev Biochem. 1995;64:29-63. doi: 10.1146/annurev.bi.64.070195.000333. Annu Rev Biochem. 1995. PMID: 7574484 Review.
-
MmpL transporter-mediated export of cell-wall associated lipids and siderophores in mycobacteria.Tuberculosis (Edinb). 2016 Sep;100:32-45. doi: 10.1016/j.tube.2016.06.004. Epub 2016 Jun 29. Tuberculosis (Edinb). 2016. PMID: 27553408 Review.
-
Evidence for the Mycobacterial Mce4 Transporter Being a Multiprotein Complex.J Bacteriol. 2021 Apr 21;203(10):e00685-20. doi: 10.1128/JB.00685-20. Print 2021 Apr 21. J Bacteriol. 2021. PMID: 33649150 Free PMC article.
-
Esx Systems and the Mycobacterial Cell Envelope: What's the Connection?J Bacteriol. 2017 Aug 8;199(17):e00131-17. doi: 10.1128/JB.00131-17. Print 2017 Sep 1. J Bacteriol. 2017. PMID: 28461452 Free PMC article. Review.
Cited by
-
The C-terminus is essential for the stability of the mycobacterial channel protein MspA.Protein Sci. 2024 Mar;33(3):e4912. doi: 10.1002/pro.4912. Protein Sci. 2024. PMID: 38358254 Free PMC article.
-
Tuberculostearic Acid Controls Mycobacterial Membrane Compartmentalization.mBio. 2023 Apr 25;14(2):e0339622. doi: 10.1128/mbio.03396-22. Epub 2023 Mar 28. mBio. 2023. PMID: 36976029 Free PMC article.
-
The Whole Is Bigger than the Sum of Its Parts: Drug Transport in the Context of Two Membranes with Active Efflux.Chem Rev. 2021 May 12;121(9):5597-5631. doi: 10.1021/acs.chemrev.0c01137. Epub 2021 Feb 17. Chem Rev. 2021. PMID: 33596653 Free PMC article.
-
Screening of novel narrow-spectrum benzofuroxan derivatives for the treatment of multidrug-resistant tuberculosis through in silico, in vitro, and in vivo approaches.Front Microbiol. 2024 Oct 11;15:1487829. doi: 10.3389/fmicb.2024.1487829. eCollection 2024. Front Microbiol. 2024. PMID: 39464394 Free PMC article.
-
Structure, Assembly, and Function of Tripartite Efflux and Type 1 Secretion Systems in Gram-Negative Bacteria.Chem Rev. 2021 May 12;121(9):5479-5596. doi: 10.1021/acs.chemrev.1c00055. Epub 2021 Apr 28. Chem Rev. 2021. PMID: 33909410 Free PMC article. Review.
References
-
- Daffé M; Etienne G, The capsule of Mycobacterium tuberculosis and its implications for pathogenicity. Tuber Lung Dis 1999, 79, 153–169. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

