Crystal structures of ORFV125 provide insight into orf virus-mediated inhibition of apoptosis
- PMID: 33175095
- PMCID: PMC7719400
- DOI: 10.1042/BCJ20200776
Crystal structures of ORFV125 provide insight into orf virus-mediated inhibition of apoptosis
Abstract
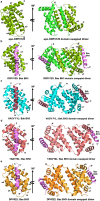
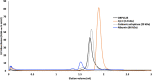
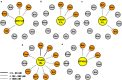
Premature apoptosis of cells is a strategy utilized by multicellular organisms to counter microbial threats. Orf virus (ORFV) is a large double-stranded DNA virus belonging to the poxviridae. ORFV encodes for an apoptosis inhibitory protein ORFV125 homologous to B-cell lymphoma 2 or Bcl-2 family proteins, which has been shown to inhibit host cell encoded pro-apoptotic Bcl-2 proteins. However, the structural basis of apoptosis inhibition by ORFV125 remains to be clarified. We show that ORFV125 is able to bind to a range of peptides spanning the BH3 motif of human pro-apoptotic Bcl-2 proteins including Bax, Bak, Puma and Hrk with modest to weak affinity. We then determined the crystal structures of ORFV125 alone as well as bound to the highest affinity ligand Bax BH3 motif. ORFV125 adopts a globular Bcl-2 fold comprising 7 α-helices, and utilizes the canonical Bcl-2 binding groove to engage pro-apoptotic host cell Bcl-2 proteins. In contrast with a previously predicted structure, ORFV125 adopts a domain-swapped dimeric topology, where the α1 helix from one protomer is swapped into a neighbouring unit. Furthermore, ORFV125 differs from the conserved architecture of the Bcl-2 binding groove and instead of α3 helix forming one of the binding groove walls, ORFV125 utilizes an extended α2 helix that comprises the equivalent region of helix α3. This results in a subtle variation of previously observed dimeric Bcl-2 architectures in other poxvirus and human encoded Bcl-2 proteins. Overall, our results provide a structural and mechanistic basis for orf virus-mediated inhibition of host cell apoptosis.
Keywords: Bcl-2; apoptosis; crystallography; orf virus; poxvirus; virology.
© 2020 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures



Similar articles
-
Structural Investigation of Orf Virus Bcl-2 Homolog ORFV125 Interactions with BH3-Motifs from BH3-Only Proteins Puma and Hrk.Viruses. 2021 Jul 15;13(7):1374. doi: 10.3390/v13071374. Viruses. 2021. PMID: 34372579 Free PMC article.
-
Structural insight into tanapoxvirus-mediated inhibition of apoptosis.FEBS J. 2020 Sep;287(17):3733-3750. doi: 10.1111/febs.15365. Epub 2020 May 31. FEBS J. 2020. PMID: 32412687
-
A novel Bcl-2-like inhibitor of apoptosis is encoded by the parapoxvirus ORF virus.J Virol. 2007 Jul;81(13):7178-88. doi: 10.1128/JVI.00404-07. Epub 2007 May 2. J Virol. 2007. PMID: 17475653 Free PMC article.
-
Structural biology of the Bcl-2 family of proteins.Biochim Biophys Acta. 2004 Mar 1;1644(2-3):83-94. doi: 10.1016/j.bbamcr.2003.08.012. Biochim Biophys Acta. 2004. PMID: 14996493 Review.
-
Structural biology of the Bcl-2 family and its mimicry by viral proteins.Cell Death Dis. 2013 Nov 7;4(11):e909. doi: 10.1038/cddis.2013.436. Cell Death Dis. 2013. PMID: 24201808 Free PMC article. Review.
Cited by
-
Contagious ecthyma in small ruminants: from etiology to vaccine challenges - a review.Vet Res Commun. 2025 Feb 24;49(2):115. doi: 10.1007/s11259-025-10677-0. Vet Res Commun. 2025. PMID: 39992468 Review.
-
Mastering Death: The Roles of Viral Bcl-2 in dsDNA Viruses.Viruses. 2024 May 30;16(6):879. doi: 10.3390/v16060879. Viruses. 2024. PMID: 38932171 Free PMC article. Review.
-
Poxviral Strategies to Overcome Host Cell Apoptosis.Pathogens. 2020 Dec 23;10(1):6. doi: 10.3390/pathogens10010006. Pathogens. 2020. PMID: 33374867 Free PMC article. Review.
-
Structural Details of BH3 Motifs and BH3-Mediated Interactions: an Updated Perspective.Front Mol Biosci. 2022 May 24;9:864874. doi: 10.3389/fmolb.2022.864874. eCollection 2022. Front Mol Biosci. 2022. PMID: 35685242 Free PMC article. Review.
-
Crystal Structures of Epstein-Barr Virus Bcl-2 Homolog BHRF1 Bound to Bid and Puma BH3 Motif Peptides.Viruses. 2022 Oct 9;14(10):2222. doi: 10.3390/v14102222. Viruses. 2022. PMID: 36298777 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

